Regulatory modules controlling maize inflorescence architecture
- PMID: 24307553
- PMCID: PMC3941108
- DOI: 10.1101/gr.166397.113
Regulatory modules controlling maize inflorescence architecture
Abstract
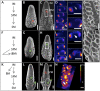
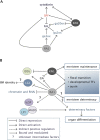
Genetic control of branching is a primary determinant of yield, regulating seed number and harvesting ability, yet little is known about the molecular networks that shape grain-bearing inflorescences of cereal crops. Here, we used the maize (Zea mays) inflorescence to investigate gene networks that modulate determinacy, specifically the decision to allow branch growth. We characterized developmental transitions by associating spatiotemporal expression profiles with morphological changes resulting from genetic perturbations that disrupt steps in a pathway controlling branching. Developmental dynamics of genes targeted in vivo by the transcription factor RAMOSA1, a key regulator of determinacy, revealed potential mechanisms for repressing branches in distinct stem cell populations, including interactions with KNOTTED1, a master regulator of stem cell maintenance. Our results uncover discrete developmental modules that function in determining grass-specific morphology and provide a basis for targeted crop improvement and translation to other cereal crops with comparable inflorescence architectures.
Figures




References
-
- Ariel FD, Manavella PA, Dezar CA, Chan RL 2007. The true story of the HD-Zip family. Trends Plant Sci 12: 419–426 - PubMed
-
- Berger N, Dubreucq B 2012. Evolution goes GAGA: GAGA binding proteins across kingdoms. Biochim Biophys Acta 1819: 863–868 - PubMed
-
- Bomblies K, Wang RL, Ambrose BA, Schmidt RJ, Meeley RB, Doebley J 2003. Duplicate FLORICAULA/LEAFY homologs zfl1 and zfl2 control inflorescence architecture and flower patterning in maize. Development 130: 2385–2395 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
