Mucosal immune responses predict clinical outcomes during influenza infection independently of age and viral load
- PMID: 24308446
- PMCID: PMC3977720
- DOI: 10.1164/rccm.201309-1616OC
Mucosal immune responses predict clinical outcomes during influenza infection independently of age and viral load
Abstract
Rationale: Children are an at-risk population for developing complications following influenza infection, but immunologic correlates of disease severity are not understood. We hypothesized that innate cellular immune responses at the site of infection would correlate with disease outcome.
Objectives: To test the immunologic basis of severe illness during natural influenza virus infection of children and adults at the site of infection.
Methods: An observational cohort study with longitudinal sampling of peripheral and mucosal sites in 84 naturally influenza-infected individuals, including infants. Cellular responses, viral loads, and cytokines were quantified from nasal lavages and blood, and correlated to clinical severity.
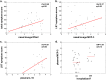
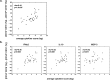
Measurements and main results: We show for the first time that although viral loads in children and adults were similar, innate responses in the airways were stronger in children and varied considerably between plasma and site of infection. Adjusting for age and viral load, an innate immune profile characterized by increased nasal lavage monocyte chemotactic protein-3, IFN-α2, and plasma IL-10 levels at enrollment predicted progression to severe disease. Increased plasma IL-10, monocyte chemotactic protein-3, and IL-6 levels predicted hospitalization. This inflammatory cytokine production correlated significantly with monocyte localization from the blood to the site of infection, with conventional monocytes positively correlating with inflammation. Increased frequencies of CD14(lo) monocytes were in the airways of participants with lower inflammatory cytokine levels.
Conclusions: An innate profile was identified that correlated with disease progression independent of viral dynamics and age. The airways and blood displayed dramatically different immune profiles emphasizing the importance of cellular migration and localized immune phenotypes.
Figures





Comment in
-
It is time for a comprehensive analytical approach in clinical studies. Virus-versus-host interaction and influenza disease severity.Am J Respir Crit Care Med. 2014 Feb 15;189(4):385-6. doi: 10.1164/rccm.201401-0008ED. Am J Respir Crit Care Med. 2014. PMID: 24528315 No abstract available.
References
-
- Liu W, de Vlas SJ, Tang F, Ma MJ, Wei MT, Liu LJ, Li ZD, Zhang L, Xin ZT, Tong YG, et al. Clinical and immunological characteristics of patients with 2009 pandemic influenza A (H1N1) virus infection after vaccination. Clin Infect Dis. 2010;51:1028–1032. - PubMed
-
- Johnson PR, Feldman S, Thompson JM, Mahoney JD, Wright PF. Immunity to influenza A virus infection in young children: a comparison of natural infection, live cold-adapted vaccine, and inactivated vaccine. J Infect Dis. 1986;154:121–127. - PubMed
-
- Richards KA, Topham D, Chaves FA, Sant AJ. Cutting edge: CD4 T cells generated from encounter with seasonal influenza viruses and vaccines have broad protein specificity and can directly recognize naturally generated epitopes derived from the live pandemic H1N1 virus. J Immunol. 2010;185:4998–5002. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

