LigSearch: a knowledge-based web server to identify likely ligands for a protein target
- PMID: 24311580
- PMCID: PMC3852652
- DOI: 10.1107/S0907444913022294
LigSearch: a knowledge-based web server to identify likely ligands for a protein target
Abstract
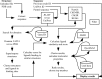
Identifying which ligands might bind to a protein before crystallization trials could provide a significant saving in time and resources. LigSearch, a web server aimed at predicting ligands that might bind to and stabilize a given protein, has been developed. Using a protein sequence and/or structure, the system searches against a variety of databases, combining available knowledge, and provides a clustered and ranked output of possible ligands. LigSearch can be accessed at http://www.ebi.ac.uk/thornton-srv/databases/LigSearch.
Keywords: LigSearch; ligand prediction.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

