TRIMmunity: the roles of the TRIM E3-ubiquitin ligase family in innate antiviral immunity
- PMID: 24333484
- PMCID: PMC3945521
- DOI: 10.1016/j.jmb.2013.12.005
TRIMmunity: the roles of the TRIM E3-ubiquitin ligase family in innate antiviral immunity
Abstract
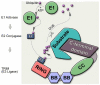
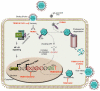
Tripartite motif (TRIM) proteins have been implicated in multiple cellular functions, including antiviral activity. Research efforts so far indicate that the antiviral activity of TRIMs relies, for the most part, on their function as E3-ubiquitin ligases. A substantial number of the TRIM family members have been demonstrated to mediate innate immune cell signal transduction and subsequent cytokine induction. In addition, a subset of TRIMs has been shown to restrict viral replication by directly targeting viral proteins. Although the body of work on the cellular roles of TRIM E3-ubiquitin ligases has rapidly grown over the last years, many aspects of their molecular workings and multi-functionality remain unclear. The antiviral function of many TRIMs seems to be conferred by specific isoforms, by sub-cellular localization and in cell-type-specific contexts. Here we review recent findings on TRIM antiviral functions, current limitations and an outlook for future research.
Keywords: E3-ubiquitin ligase; antiviral response; innate immunity; restriction factors; tripartite motif.
Copyright © 2013 Elsevier Ltd. All rights reserved.
Figures





Similar articles
-
To TRIM or not to TRIM: the balance of host-virus interactions mediated by the ubiquitin system.J Gen Virol. 2019 Dec;100(12):1641-1662. doi: 10.1099/jgv.0.001341. J Gen Virol. 2019. PMID: 31661051 Free PMC article.
-
The Roles of TRIMs in Antiviral Innate Immune Signaling.Front Cell Infect Microbiol. 2021 Mar 15;11:628275. doi: 10.3389/fcimb.2021.628275. eCollection 2021. Front Cell Infect Microbiol. 2021. PMID: 33791238 Free PMC article. Review.
-
TRIM Proteins and Their Roles in Antiviral Host Defenses.Annu Rev Virol. 2018 Sep 29;5(1):385-405. doi: 10.1146/annurev-virology-092917-043323. Epub 2018 Jun 27. Annu Rev Virol. 2018. PMID: 29949725 Free PMC article. Review.
-
TRIM proteins: A 'swiss army knife' of antiviral immunity.PLoS Pathog. 2025 May 12;21(5):e1013147. doi: 10.1371/journal.ppat.1013147. eCollection 2025 May. PLoS Pathog. 2025. PMID: 40354393 Free PMC article. Review.
-
The TRIMendous Role of TRIMs in Virus-Host Interactions.Vaccines (Basel). 2017 Aug 22;5(3):23. doi: 10.3390/vaccines5030023. Vaccines (Basel). 2017. PMID: 28829373 Free PMC article. Review.
Cited by
-
The Transcriptional Changes of trim Genes Associated with Parkinson's Disease on a Model of Human Induced Pluripotent Stem Cells.Mol Neurobiol. 2017 Nov;54(9):7204-7211. doi: 10.1007/s12035-016-0230-7. Epub 2016 Oct 29. Mol Neurobiol. 2017. PMID: 27796756
-
TRIM56 restricts Coxsackievirus B infection by mediating the ubiquitination of viral RNA-dependent RNA polymerase 3D.PLoS Pathog. 2024 Sep 30;20(9):e1012594. doi: 10.1371/journal.ppat.1012594. eCollection 2024 Sep. PLoS Pathog. 2024. PMID: 39348396 Free PMC article.
-
TRIM34 localizes to the mitochondria and mediates apoptosis through the mitochondrial pathway in HEK293T cells.Heliyon. 2020 Jan 9;6(1):e03115. doi: 10.1016/j.heliyon.2019.e03115. eCollection 2020 Jan. Heliyon. 2020. PMID: 31956709 Free PMC article.
-
TRIM25 inhibits infectious bursal disease virus replication by targeting VP3 for ubiquitination and degradation.PLoS Pathog. 2021 Sep 13;17(9):e1009900. doi: 10.1371/journal.ppat.1009900. eCollection 2021 Sep. PLoS Pathog. 2021. PMID: 34516573 Free PMC article.
-
Swine acute diarrhea syndrome coronavirus nucleocapsid protein antagonizes the IFN response through inhibiting TRIM25 oligomerization and functional activation of RIG-I/TRIM25.Vet Res. 2024 Apr 8;55(1):44. doi: 10.1186/s13567-024-01303-z. Vet Res. 2024. PMID: 38589930 Free PMC article.
References
-
- Medzhitov R. Recognition of microorganisms and activation of the immune response. Nature. 2007;449:819–26. - PubMed
-
- Bieniasz PD. Intrinsic immunity: a front-line defense against viral attack. Nat Immunol. 2004;5:1109–15. - PubMed
-
- Platanias LC. Mechanisms of type-I- and type-II-interferon-mediated signalling. Nat Rev Immunol. 2005;5:375–86. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P01AI090935/AI/NIAID NIH HHS/United States
- U54AI057158/AI/NIAID NIH HHS/United States
- R01DA033773/DA/NIDA NIH HHS/United States
- U19AI106754/AI/NIAID NIH HHS/United States
- U54 AI057158/AI/NIAID NIH HHS/United States
- U01 AI095611/AI/NIAID NIH HHS/United States
- U19AI083025/AI/NIAID NIH HHS/United States
- R01 DA033773/DA/NIDA NIH HHS/United States
- HHSN266200700010C/AI/NIAID NIH HHS/United States
- U19AI089987/AI/NIAID NIH HHS/United States
- R01 AI046954/AI/NIAID NIH HHS/United States
- HHSN272201000054C/AI/NIAID NIH HHS/United States
- U01AI095611/AI/NIAID NIH HHS/United States
- U19 AI083025/AI/NIAID NIH HHS/United States
- R01AI046954/AI/NIAID NIH HHS/United States
- U19 AI106754/AI/NIAID NIH HHS/United States
- U19 AI089987/AI/NIAID NIH HHS/United States
- P01 AI090935/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

