Hairless is a histone H3K9 demethylase
- PMID: 24334705
- PMCID: PMC3963012
- DOI: 10.1096/fj.13-237677
Hairless is a histone H3K9 demethylase
Abstract
The hairless (HR) protein contains a Jumonji C (JmjC) domain that is conserved among a family of proteins with histone demethylase (HDM) activity. To test whether HR possesses HDM activity, we performed a series of in vitro demethylation assays, which demonstrated that HR can demethylate monomethylated or dimethylated histone H3 lysine 9 (H3K9me1 or me2). Moreover, ectopic expression of wild-type HR, but not JmjC-mutant HR, led to pronounced demethylation of H3K9 in cultured human HeLa cells. We also show that two missense mutations in HR, which we and others described in patients with atrichia with papular lesions, abolished the demethylase activity of HR, demonstrating the role of HR demethylase activity in human disease. By ChIP-Seq analysis, we identified multiple new HR target genes, many of which play important roles in epidermal development, neural function, and transcriptional regulation, consistent with the predicted biological functions of HR. Our findings demonstrate for the first time that HR is a H3K9 demethylase that regulates epidermal homeostasis via direct control of its target genes.
Keywords: ChIP-Seq; epigenetics.
Figures


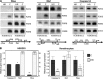
References
-
- Peters A. H., O'Carroll D., Scherthan H., Mechtler K., Sauer S., Schofer C., Weipoltshammer K., Pagani M., Lachner M., Kohlmaier A., Opravil S., Doyle M., Sibilia M., Jenuwein T. (2001) Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107, 323–337 - PubMed
-
- Tachibana M., Sugimoto K., Nozaki M., Ueda J., Ohta T., Ohki M., Fukuda M., Takeda N., Niida H., Kato H., Shinkai Y. (2002) G9a histone methyltransferase plays a dominant role in euchromatic histone H3 lysine 9 methylation and is essential for early embryogenesis. Genes Dev. 16, 1779–1791 - PMC - PubMed
-
- Bernstein B. E., Mikkelsen T. S., Xie X., Kamal M., Huebert D. J., Cuff J., Fry B., Meissner A., Wernig M., Plath K., Jaenisch R., Wagschal A., Feil R., Schreiber S. L., Lander E. S. (2006) A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 - PubMed
-
- Agger K., Cloos P. A., Christensen J., Pasini D., Rose S., Rappsilber J., Issaeva I., Canaani E., Salcini A. E., Helin K. (2007) UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 449, 731–734 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

