Structure of the mammalian ribosomal pre-termination complex associated with eRF1.eRF3.GDPNP
- PMID: 24335085
- PMCID: PMC3950680
- DOI: 10.1093/nar/gkt1279
Structure of the mammalian ribosomal pre-termination complex associated with eRF1.eRF3.GDPNP
Abstract
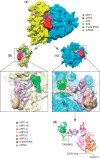
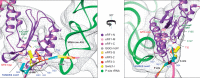
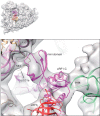
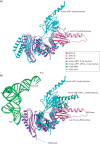
Eukaryotic translation termination results from the complex functional interplay between two release factors, eRF1 and eRF3, in which GTP hydrolysis by eRF3 couples codon recognition with peptidyl-tRNA hydrolysis by eRF1. Here, we present a cryo-electron microscopy structure of pre-termination complexes associated with eRF1•eRF3•GDPNP at 9.7 -Å resolution, which corresponds to the initial pre-GTP hydrolysis stage of factor attachment and stop codon recognition. It reveals the ribosomal positions of eRFs and provides insights into the mechanisms of stop codon recognition and triggering of eRF3's GTPase activity.
Figures


References
-
- Jackson RJ, Hellen CU, Pestova TV. Termination and post-termination events in eukaryotic translation. Adv. Protein Chem. Struct. Biol. 2012;86:45–93. - PubMed
-
- Alkalaeva EZ, Pisarev AV, Frolova LY, Kisselev LL, Pestova TV. In vitro reconstitution of eukaryotic translation reveals cooperativity between release factors eRF1 and eRF3. Cell. 2006;125:1125–1136. - PubMed
-
- Pisareva VP, Pisarev AV, Hellen CU, Rodnina MV, Pestova TV. Kinetic analysis of interaction of eukaryotic release factor 3 with guanine nucleotides. J. Biol. Chem. 2006;281:40224–40235. - PubMed
-
- Mitkevich VA, Kononenko AV, Petrushanko IY, Yanvarev DV, Makarov AA, Kisselev LL. Termination of translation in eukaryotes is mediated by the quaternary eRF1*eRF3*GTP*Mg2+ complex. The biological roles of eRF3 and prokaryotic RF3 are profoundly distinct. Nucleic Acids Res. 2008;34:3947–3954. - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

