Genome-wide dissection of the quorum sensing signalling pathway in Trypanosoma brucei
- PMID: 24336212
- PMCID: PMC3908871
- DOI: 10.1038/nature12864
Genome-wide dissection of the quorum sensing signalling pathway in Trypanosoma brucei
Abstract
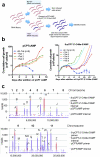
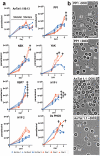
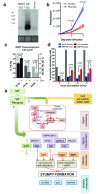
The protozoan parasites Trypanosoma brucei spp. cause important human and livestock diseases in sub-Saharan Africa. In mammalian blood, two developmental forms of the parasite exist: proliferative 'slender' forms and arrested 'stumpy' forms that are responsible for transmission to tsetse flies. The slender to stumpy differentiation is a density-dependent response that resembles quorum sensing in microbial systems and is crucial for the parasite life cycle, ensuring both infection chronicity and disease transmission. This response is triggered by an elusive 'stumpy induction factor' (SIF) whose intracellular signalling pathway is also uncharacterized. Laboratory-adapted (monomorphic) trypanosome strains respond inefficiently to SIF but can generate forms with stumpy characteristics when exposed to cell-permeable cAMP and AMP analogues. Exploiting this, we have used a genome-wide RNA interference library screen to identify the signalling components driving stumpy formation. In separate screens, monomorphic parasites were exposed to 8-(4-chlorophenylthio)-cAMP (pCPT-cAMP) or 8-pCPT-2'-O-methyl-5'-AMP to select cells that were unresponsive to these signals and hence remained proliferative. Genome-wide Ion Torrent based RNAi target sequencing identified cohorts of genes implicated in each step of the signalling pathway, from purine metabolism, through signal transducers (kinases, phosphatases) to gene expression regulators. Genes at each step were independently validated in cells naturally capable of stumpy formation, confirming their role in density sensing in vivo. The putative RNA-binding protein, RBP7, was required for normal quorum sensing and promoted cell-cycle arrest and transmission competence when overexpressed. This study reveals that quorum sensing signalling in trypanosomes shares similarities to fundamental quiescence pathways in eukaryotic cells, its components providing targets for quorum-sensing interference-based therapeutics.
Figures

References
-
- Goldenberg S, Avila AR. Aspects of Trypanosoma cruzi stage differentiation. Advances in parasitology. 2011;75:285–305. doi:10.1016/B978-0-12-385863-4.00013-7B978-0-12-385863-4.00013-7 [pii] - PubMed
Methods References
-
- Redmond S, Vadivelu J, Field MC. RNAit: an automated web-based tool for the selection of RNAi targets in Trypanosoma brucei. Mol Biochem Parasitol. 2003;128:115–118. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

