OnPLS integration of transcriptomic, proteomic and metabolomic data shows multi-level oxidative stress responses in the cambium of transgenic hipI- superoxide dismutase Populus plants
- PMID: 24341908
- PMCID: PMC3878592
- DOI: 10.1186/1471-2164-14-893
OnPLS integration of transcriptomic, proteomic and metabolomic data shows multi-level oxidative stress responses in the cambium of transgenic hipI- superoxide dismutase Populus plants
Abstract
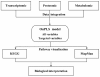
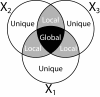
Background: Reactive oxygen species (ROS) are involved in the regulation of diverse physiological processes in plants, including various biotic and abiotic stress responses. Thus, oxidative stress tolerance mechanisms in plants are complex, and diverse responses at multiple levels need to be characterized in order to understand them. Here we present system responses to oxidative stress in Populus by integrating data from analyses of the cambial region of wild-type controls and plants expressing high-isoelectric-point superoxide dismutase (hipI-SOD) transcripts in antisense orientation showing a higher production of superoxide. The cambium, a thin cell layer, generates cells that differentiate to form either phloem or xylem and is hypothesized to be a major reason for phenotypic perturbations in the transgenic plants. Data from multiple platforms including transcriptomics (microarray analysis), proteomics (UPLC/QTOF-MS), and metabolomics (GC-TOF/MS, UPLC/MS, and UHPLC-LTQ/MS) were integrated using the most recent development of orthogonal projections to latent structures called OnPLS. OnPLS is a symmetrical multi-block method that does not depend on the order of analysis when more than two blocks are analysed. Significantly affected genes, proteins and metabolites were then visualized in painted pathway diagrams.
Results: The main categories that appear to be significantly influenced in the transgenic plants were pathways related to redox regulation, carbon metabolism and protein degradation, e.g. the glycolysis and pentose phosphate pathways (PPP). The results provide system-level information on ROS metabolism and responses to oxidative stress, and indicate that some initial responses to oxidative stress may share common pathways.
Conclusion: The proposed data evaluation strategy shows an efficient way of compiling complex, multi-platform datasets to obtain significant biological information.
Figures


References
-
- Higashi Y, Hirai MY, Fujiwara T, Naito S, Noji M, Saito K. Proteomic and transcriptomic analysis of Arabidopsis seeds: molecular evidence for successive processing of seed proteins and its implication in the stress response to sulfur nutrition. Plant J. 2006;14:557–571. doi: 10.1111/j.1365-313X.2006.02900.x. - DOI - PubMed
-
- Hirai MY, Yano M, Goodenowe DB, Kanaya S, Kimura T, Awazuhara M, Arita M, Fujiwara T, Saito K. Integration of transcriptomics and metabolomics for understanding of global responses to nutritional stresses in Arabidopsis thaliana. Proc Natl Acad Sci U S A. 2004;14:10205–10210. doi: 10.1073/pnas.0403218101. - DOI - PMC - PubMed
-
- Hirai MY, Klein M, Fujikawa Y, Yano M, Goodenowe DB, Yamazaki Y, Kanaya S, Nakamura Y, Kitayama M, Suzuki H, Sakurai N, Shibata D, Tokuhisa J, Reichelt M, Gershenzon J, Papenbrock J, Saito K. Elucidation of gene-to-gene and metabolite-to-gene networks in Arabidopsis by integration of metabolomics and transcriptomics. J Biol Chem. 2005;14:25590–25595. doi: 10.1074/jbc.M502332200. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

