How do bacteria localize proteins to the cell pole?
- PMID: 24345373
- PMCID: PMC3874780
- DOI: 10.1242/jcs.138628
How do bacteria localize proteins to the cell pole?
Abstract
It is now well appreciated that bacterial cells are highly organized, which is far from the initial concept that they are merely bags of randomly distributed macromolecules and chemicals. Central to their spatial organization is the precise positioning of certain proteins in subcellular domains of the cell. In particular, the cell poles - the ends of rod-shaped cells - constitute important platforms for cellular regulation that underlie processes as essential as cell cycle progression, cellular differentiation, virulence, chemotaxis and growth of appendages. Thus, understanding how the polar localization of specific proteins is achieved and regulated is a crucial question in bacterial cell biology. Often, polarly localized proteins are recruited to the poles through their interaction with other proteins or protein complexes that were already located there, in a so-called diffusion-and-capture mechanism. Bacteria are also starting to reveal their secrets on how the initial pole 'recognition' can occur and how this event can be regulated to generate dynamic, reproducible patterns in time (for example, during the cell cycle) and space (for example, at a specific cell pole). Here, we review the major mechanisms that have been described in the literature, with an emphasis on the self-organizing principles. We also present regulation strategies adopted by bacterial cells to obtain complex spatiotemporal patterns of protein localization.
Keywords: Bacterial cell cycle; Polar localization; Spatial organization.
Figures


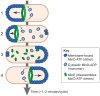

References
-
- Bowman G. R., Comolli L. R., Gaietta G. M., Fero M., Hong S-H., Jones Y., Lee J. H., Downing K. H., Ellisman M. H., McAdams H. H. et al. (2010). Caulobacter PopZ forms a polar subdomain dictating sequential changes in pole composition and function. Mol. Microbiol. 76, 173–189 10.1111/j.1365-2958.2010.07088.x - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

