A domesticated PiggyBac transposase interacts with heterochromatin and catalyzes reproducible DNA elimination in Tetrahymena
- PMID: 24348275
- PMCID: PMC3861120
- DOI: 10.1371/journal.pgen.1004032
A domesticated PiggyBac transposase interacts with heterochromatin and catalyzes reproducible DNA elimination in Tetrahymena
Abstract
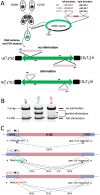
The somatic genome of the ciliated protist Tetrahymena undergoes DNA elimination of defined sequences called internal eliminated sequences (IESs), which account for ~30% of the germline genome. During DNA elimination, IES regions are heterochromatinized and assembled into heterochromatin bodies in the developing somatic nucleus. The domesticated piggyBac transposase Tpb2p is essential for the formation of heterochromatin bodies and DNA elimination. In this study, we demonstrate that the activities of Tpb2p involved in forming heterochromatin bodies and executing DNA elimination are genetically separable. The cysteine-rich domain of Tpb2p, which interacts with the heterochromatin-specific histone modifications, is necessary for both heterochromatin body formation and DNA elimination, whereas the endonuclease activity of Tpb2p is only necessary for DNA elimination. Furthermore, we demonstrate that the endonuclease activity of Tpb2p in vitro and the endonuclease activity that executes DNA elimination in vivo have similar substrate sequence preferences. These results strongly indicate that Tpb2p is the endonuclease that directly catalyzes the excision of IESs and that the boundaries of IESs are at least partially determined by the combination of Tpb2p-heterochromatin interaction and relaxed sequence preference of the endonuclease activity of Tpb2p.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

Similar articles
-
A domesticated piggyBac transposase plays key roles in heterochromatin dynamics and DNA cleavage during programmed DNA deletion in Tetrahymena thermophila.Mol Biol Cell. 2010 May 15;21(10):1753-62. doi: 10.1091/mbc.e09-12-1079. Epub 2010 Mar 31. Mol Biol Cell. 2010. PMID: 20357003 Free PMC article.
-
DNA elimination in ciliates: transposon domestication and genome surveillance.Annu Rev Genet. 2011;45:227-46. doi: 10.1146/annurev-genet-110410-132432. Epub 2011 Sep 9. Annu Rev Genet. 2011. PMID: 21910632 Review.
-
The piggyBac transposon-derived genes TPB1 and TPB6 mediate essential transposon-like excision during the developmental rearrangement of key genes in Tetrahymena thermophila.Genes Dev. 2016 Dec 15;30(24):2724-2736. doi: 10.1101/gad.290460.116. Genes Dev. 2016. PMID: 28087716 Free PMC article.
-
The taming of the shrew: Regulation of a catalytically active domesticated transposase.Mob Genet Elements. 2014 May 27;4:e29383. doi: 10.4161/mge.29383. eCollection 2014. Mob Genet Elements. 2014. PMID: 25054083 Free PMC article.
-
Transposon domestication versus mutualism in ciliate genome rearrangements.PLoS Genet. 2013;9(8):e1003659. doi: 10.1371/journal.pgen.1003659. Epub 2013 Aug 1. PLoS Genet. 2013. PMID: 23935529 Free PMC article. Review.
Cited by
-
Biogeography and Character Evolution of the Ciliate Genus Euplotes (Spirotrichea, Euplotia), with Description of Euplotes curdsi sp. nov.PLoS One. 2016 Nov 9;11(11):e0165442. doi: 10.1371/journal.pone.0165442. eCollection 2016. PLoS One. 2016. PMID: 27828996 Free PMC article.
-
Origins of genome-editing excisases as illuminated by the somatic genome of the ciliate Blepharisma.Proc Natl Acad Sci U S A. 2023 Jan 24;120(4):e2213887120. doi: 10.1073/pnas.2213887120. Epub 2023 Jan 20. Proc Natl Acad Sci U S A. 2023. PMID: 36669098 Free PMC article.
-
MITE infestation accommodated by genome editing in the germline genome of the ciliate Blepharisma.Proc Natl Acad Sci U S A. 2023 Jan 24;120(4):e2213985120. doi: 10.1073/pnas.2213985120. Epub 2023 Jan 20. Proc Natl Acad Sci U S A. 2023. PMID: 36669106 Free PMC article.
-
The unusual structure of the PiggyMac cysteine-rich domain reveals zinc finger diversity in PiggyBac-related transposases.Mob DNA. 2021 Apr 29;12(1):12. doi: 10.1186/s13100-021-00240-4. Mob DNA. 2021. PMID: 33926516 Free PMC article.
-
Constitutive Heterochromatin in Eukaryotic Genomes: A Mine of Transposable Elements.Cells. 2022 Feb 22;11(5):761. doi: 10.3390/cells11050761. Cells. 2022. PMID: 35269383 Free PMC article. Review.
References
-
- Orgel LE, Crick FH (1980) Selfish DNA: the ultimate parasite. Nature 284: 604–607. - PubMed
-
- Almeida R, Allshire RC (2005) RNA silencing and genome regulation. Trends Cell Biol 15: 251–258 doi:10.1016/j.tcb.2005.03.006 - DOI - PubMed
-
- Volff J-N (2006) Turning junk into gold: domestication of transposable elements and the creation of new genes in eukaryotes. Bioessays 28: 913–922 doi:10.1002/bies.20452 - DOI - PubMed
-
- Cheng C-Y, Vogt A, Mochizuki K, Yao M-C (2010) A domesticated piggyBac transposase plays key roles in heterochromatin dynamics and DNA cleavage during programmed DNA deletion in Tetrahymena thermophila. Mol Biol Cell 21: 1753–1762 doi:10.1091/mbc.E09-12-1079 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

