Endosomal transport of septin mRNA and protein indicates local translation on endosomes and is required for correct septin filamentation
- PMID: 24355572
- PMCID: PMC4303453
- DOI: 10.1002/embr.201338037
Endosomal transport of septin mRNA and protein indicates local translation on endosomes and is required for correct septin filamentation
Abstract
Endosomes transport lipids and proteins over long distances by shuttling along microtubules. They also carry mRNAs on their surface, but the precise molecular function of this trafficking process is unknown. By live cell imaging of polarized fungal hyphae, we show microtubule-dependent transport of septin mRNA and encoded septin protein on the same shuttling endosomes. Consistent with the hypothesis that septin mRNA is translated on endosomes, the accumulation of septin protein on endosomes requires the recruitment of septin mRNA. Furthermore, ribosomal proteins co-localise with shuttling endosomes, but only if mRNA is present. Importantly, endosomal trafficking is essential for an efficient delivery of septin protein to filaments at growth poles, a process necessary to establish unipolar growth. Thus, we propose that local mRNA translation loads endosomes with septins for assembly and efficient delivery to septin filaments.
Figures
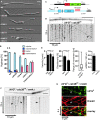
Hyphae of AB33 derivatives 4 h after induction of polar growth (DIC images; size bar, 10 μm).
Percentages of hyphae after 4, 6 and 9 h p.i.. Unipolarity, bipolarity and septum formation was quantified (error bars, s.e.m.; n = 3 independent experiments, 2422 filaments were counted in total; note that septum formation is given relative to the values of unipolar or bipolar hyphae set to 100%).
Components of the modified λN* RNA reporter system (Pcrg1, arabinose-regulated promoter; Tnos, heterologous transcriptional terminator; cdc3B16 carries 16 copies of boxB hairpin in its 3′ UTR).
Hyphal tip of a strain expressing the λN*G2 protein and cdc3B16 mRNA. Micrograph (size bar, 10 μm) and the corresponding kymograph show directed particles (arrowheads; supplementary Movie S3).
Velocity (total directed particles and those moving towards growth pole or septum; error bars represent s.e.m.; one way ANOVA, P > 0.05); directionality of particles (error bars represent s.e.m.; paired two-tailed t-test, P > 0.05); range of directed movement (whisker diagram showing median, min/max values and 25/75 percentiles); number of directed mRNPs in 100 μm of hyphae (n = 3, 69 particles in 97 wt hyphae and 0 particles in 37 rrm4Δ hyphae; note, length of hyphae varies in tested strains; error bars represent s.e.m.).
Micrograph (size bar, 10 μm) and kymographs (bottom) showing restricted moving and static particles (arrowheads and arrow, respectively; supplementary Movie S4).
Kymographs of λN*G2-labelled cdc3B16 mRNA (green) and Rrm4C (red). Co-localisation (yellow) is indicated by arrowheads.

Micrographs of Cdc3G-expressing hyphae (z-plane 0 or maximum projection of z-stacks with 0.25 μm steps; size bar, 10 μm). Hypha at the bottom was treated with benomyl for 1 h (arrowheads mark small circle; see inlay, size bar, 1 μm).
Bipolarly growing cdc3G/rrm4Δ hypha (size bars 10, 1 and 5 μm for top, bottom left and bottom right, respectively).
Kymographs of Cdc3G expressing hypha (asterisk marks septum; top untreated and bottom treated with benomyl).
Micrograph (top) and corresponding kymograph (bottom) of cdc3G/rrm4Δ hypha (size bar, 10 μm; supplementary Movie S7).
Kymographs showing co-localisation of Cdc3G and Rrm4C (arrowheads).

A Top: schematic representation of the construct expressing Cdc3CB16 (constitutively active Potef promoter; Cdc3 fused to mCherry and 16 boxB binding sites in the 3′UTR). Bottom: kymographs of movies recorded in strain expressing Cdc3CB16. Arrowheads indicate co-localising signals.
B Micrographs and corresponding kymographs depicting bidirectional movement of Rrm4C and Pab1G.
C Movement of Pab1G and Rrm4RmR1 fused to mRfp shown as in panel (B). Arrowheads indicate residual movement of Pab1G.
D, E Kymographs depicting λN*G2-labelled cdc3B16 mRNA (D) or Cdc3G protein (E). Strains carry the allele rrm4RmR1. Arrowheads highlight movement of cdc3B16 mRNAs
F Movement of Pab1G and Rrm4RRRMΔ (Rrm4 with N-terminal deletion of three RRMs fused to mRfp) shown as in panel (B).
G, H Kymographs depicting λN*G2-labelled cdc3B16 mRNA (G) or Cdc3G protein (H). Strains carry the allele rrm4RRRMΔ. Arrowheads as in panel (D).
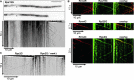
Top: micrograph before and after (upper and lower micrograph) photobleaching of Gfp fluorescence (size bar, 10 μm; asterisk indicates nucleus). Bottom: kymograph of supplementary Movie S9 (after photobleaching).
Co-localisation (yellow) of Rrm4 (red) and ribosomal proteins (green) on shuttling endosomes.
Shuttling of Rps2G is Rrm4-dependent.

A Representative FRAP experiment analysing hypha expressing Cdc3G (supplementary Movie S10; asterisk marks septum).
B, C FRAP analysis of Cdc3G at hyphal tips of indicated strains (data were fitted to uniphasic exponential equation, dotted lines indicate half time of recovery; n = 6 and n = 3 in (B) and (C), respectively; error bars represent s.e.m.). Fluorescence is normalised to plateau, further details in supplementary Fig S4.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

