Regulatory RNAs: charming gene management styles for synthetic biology applications
- PMID: 24356572
- PMCID: PMC3917981
- DOI: 10.4161/rna.27102
Regulatory RNAs: charming gene management styles for synthetic biology applications
Abstract
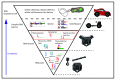
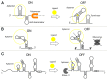
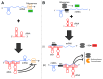
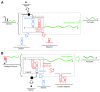
RNAs have many important functional properties, including that they are independently controllable and highly tunable. As a result of these advantageous properties, their use in a myriad of sophisticated devices has been widely explored. Yet, the exploitation of RNAs for synthetic applications is highly dependent on the ability to characterize the many new molecules that continue to be discovered by large-scale sequencing and high-throughput screening techniques. In this review, we present an exhaustive survey of the most recent synthetic bacterial riboswitches and small RNAs while emphasizing their virtues in gene expression management. We also explore the use of these RNA components as building blocks in the RNA synthetic biology toolbox and discuss examples of synthetic RNA components used to rewire bacterial regulatory circuitry. We anticipate that this field will expand its catalog of smart devices by mimicking and manipulating natural RNA mechanisms and functions.
Keywords: RNA regulation; RNAs and biotechnology; riboswitches; small RNAs; synthetic RNAs; transcriptional control; translational control.
Figures
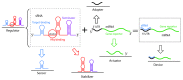
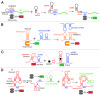
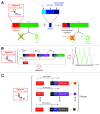
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
