Inhibitory effects of caffeic acid phenethyl ester derivatives on replication of hepatitis C virus
- PMID: 24358168
- PMCID: PMC3866116
- DOI: 10.1371/journal.pone.0082299
Inhibitory effects of caffeic acid phenethyl ester derivatives on replication of hepatitis C virus
Abstract
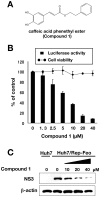
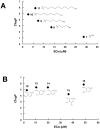
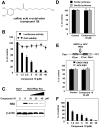
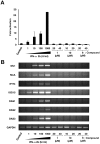
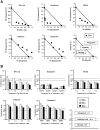
Caffeic acid phenethyl ester (CAPE) has been reported as a multifunctional compound. In this report, we tested the effect of CAPE and its derivatives on hepatitis C virus (HCV) replication in order to develop an effective anti-HCV compound. CAPE and CAPE derivatives exhibited anti-HCV activity against an HCV replicon cell line of genotype 1b with EC50 values in a range from 1.0 to 109.6 µM. Analyses of chemical structure and antiviral activity suggested that the length of the n-alkyl side chain and catechol moiety are responsible for the anti-HCV activity of these compounds. Caffeic acid n-octyl ester exhibited the highest anti-HCV activity among the tested derivatives with an EC50 value of 1.0 µM and an SI value of 63.1 by using the replicon cell line derived from genotype 1b strain Con1. Treatment with caffeic acid n-octyl ester inhibited HCV replication of genotype 2a at a similar level to that of genotype 1b irrespectively of interferon signaling. Caffeic acid n-octyl ester could synergistically enhance the anti-HCV activities of interferon-alpha 2b, daclatasvir, and VX-222, but neither telaprevir nor danoprevir. These results suggest that caffeic acid n-octyl ester is a potential candidate for novel anti-HCV chemotherapy drugs.
Conflict of interest statement
Figures
References
-
- Baldo V, Baldovin T, Trivello R, Floreani A (2008) Epidemiology of HCV infection. Curr Pharm Des 14: 1646–1654. - PubMed
-
- Lohmann V, Korner F, Koch J, Herian U, Theilmann L, et al. (1999) Replication of subgenomic hepatitis C virus RNAs in a hepatoma cell line. Science 285: 110–113. - PubMed
-
- Hofmann WP, Zeuzem S (2011) A new standard of care for the treatment of chronic HCV infection. Nat Rev Gastroenterol Hepatol 8: 257–264. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

