Soluble expression of human leukemia inhibitory factor with protein disulfide isomerase in Escherichia coli and its simple purification
- PMID: 24358310
- PMCID: PMC3865251
- DOI: 10.1371/journal.pone.0083781
Soluble expression of human leukemia inhibitory factor with protein disulfide isomerase in Escherichia coli and its simple purification
Erratum in
- PLoS One. 2014;9(1). doi:10.1371/annotation/4b2acdb5-5bab-4bca-8fbc-f9f293b38ee0. Jung, A Song [corrected to Song, Jung-A]
Abstract
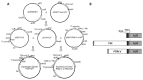
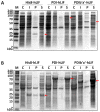
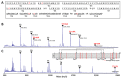
Human leukemia inhibitory factor (hLIF) is a multifunctional cytokine that is essential for maintaining the pluripotency of embryonic stem cells. hLIF may be also be useful in aiding fertility through its effects on increasing the implantation rate of fertilized eggs. Thus these applications in biomedical research and clinical medicine create a high demand for bioactive hLIF. However, production of active hLIF is problematic since eukaryotic cells demonstrate limited expression and prokaryotic cells produce insoluble protein. Here, we have adopted a hybrid protein disulfide isomerase design to increase the solubility of hLIF in Escherichia coli. Low temperature expression of hLIF fused to the b'a' domain of protein disulfide isomerase (PDIb'a') increased the soluble expression in comparison to controls. A simple purification protocol for bioactive hLIF was established that includes removal of the PDIb'a' domain by cleavage by TEV protease. The resulting hLIF, which contains one extra glycine residue at the N-terminus, was highly pure and demonstrated endotoxin levels below 0.05 EU/μg. The presence of an intramolecular disulfide bond was identified using mass spectroscopy. This purified hLIF effectively maintained the pluripotency of a murine embryonic stem cell line. Thus we have developed an effective method to produce a pure bioactive version of hLIF in E. coli for use in biomedical research.
Conflict of interest statement
Figures


Similar articles
-
Prokaryotic soluble overexpression and purification of bioactive human growth hormone by fusion to thioredoxin, maltose binding protein, and protein disulfide isomerase.PLoS One. 2014 Mar 10;9(3):e89038. doi: 10.1371/journal.pone.0089038. eCollection 2014. PLoS One. 2014. PMID: 24614134 Free PMC article.
-
Efficient production of human interleukin-3 from Escherichia coli using protein disulfide isomerase b'a' domain.Biotechnol J. 2024 May;19(5):e2300581. doi: 10.1002/biot.202300581. Biotechnol J. 2024. PMID: 38719587
-
Preparation of bioactive soluble human leukemia inhibitory factor from recombinant Escherichia coli using thioredoxin as fusion partner.Protein Expr Purif. 2010 Sep;73(1):51-7. doi: 10.1016/j.pep.2010.04.002. Epub 2010 Apr 8. Protein Expr Purif. 2010. PMID: 20381622
-
Soluble prokaryotic overexpression and purification of bioactive human granulocyte colony-stimulating factor by maltose binding protein and protein disulfide isomerase.PLoS One. 2014 Mar 3;9(3):e89906. doi: 10.1371/journal.pone.0089906. eCollection 2014. PLoS One. 2014. PMID: 24594699 Free PMC article.
-
Targeted expression, purification, and cleavage of fusion proteins from inclusion bodies in Escherichia coli.FEBS Lett. 2014 Jan 21;588(2):247-52. doi: 10.1016/j.febslet.2013.09.028. Epub 2013 Sep 27. FEBS Lett. 2014. PMID: 24076468 Review.
Cited by
-
Challenges Associated With the Formation of Recombinant Protein Inclusion Bodies in Escherichia coli and Strategies to Address Them for Industrial Applications.Front Bioeng Biotechnol. 2021 Feb 10;9:630551. doi: 10.3389/fbioe.2021.630551. eCollection 2021. Front Bioeng Biotechnol. 2021. PMID: 33644021 Free PMC article. Review.
-
Prokaryotic Soluble Overexpression and Purification of Human VEGF165 by Fusion to a Maltose Binding Protein Tag.PLoS One. 2016 May 27;11(5):e0156296. doi: 10.1371/journal.pone.0156296. eCollection 2016. PLoS One. 2016. PMID: 27231876 Free PMC article.
-
Prokaryotic soluble overexpression and purification of bioactive human growth hormone by fusion to thioredoxin, maltose binding protein, and protein disulfide isomerase.PLoS One. 2014 Mar 10;9(3):e89038. doi: 10.1371/journal.pone.0089038. eCollection 2014. PLoS One. 2014. PMID: 24614134 Free PMC article.
-
Prokaryotic soluble overexpression and purification of oncostatin M using a fusion approach and genetically engineered E. coli strains.Sci Rep. 2019 Sep 23;9(1):13706. doi: 10.1038/s41598-019-50110-6. Sci Rep. 2019. PMID: 31548569 Free PMC article.
-
Soluble Cytoplasmic Expression and Purification of Immunotoxin HER2(scFv)-PE24B as a Maltose Binding Protein Fusion.Int J Mol Sci. 2021 Jun 17;22(12):6483. doi: 10.3390/ijms22126483. Int J Mol Sci. 2021. PMID: 34204265 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

