Fgf3 and Fgf10a work in concert to promote maturation of the epibranchial placodes in zebrafish
- PMID: 24358375
- PMCID: PMC3866233
- DOI: 10.1371/journal.pone.0085087
Fgf3 and Fgf10a work in concert to promote maturation of the epibranchial placodes in zebrafish
Abstract
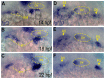
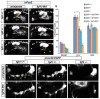
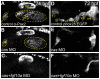
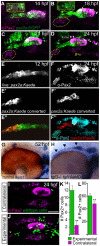
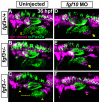
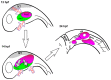
Essential cellular components of the paired sensory organs of the vertebrate head are derived from transient thickenings of embryonic ectoderm known as cranial placodes. The epibranchial (EB) placodes give rise to sensory neurons of the EB ganglia that are responsible for relaying visceral sensations form the periphery to the central nervous system. Development of EB placodes and subsequent formation of EB ganglia is a multistep process regulated by various extrinsic factors, including fibroblast growth factors (Fgfs). We discovered that two Fgf ligands, Fgf3 and Fgf10a, cooperate to promote EB placode development. Whereas EB placodes are induced in the absence of Fgf3 and Fgf10a, they fail to express placode specific markers Pax2a and Sox3. Expression analysis and mosaic rescue experiments demonstrate that Fgf3 signal is derived from the endoderm, whereas Fgf10a is emitted from the lateral line system and the otic placode. Further analyses revealed that Fgf3 and Fgf10a activities are not required for cell proliferation or survival, but are required for placodal cells to undergo neurogenesis. Based on these data, we conclude that a combined loss of these Fgf factors results in a failure of the EB placode precursors to initiate a transcriptional program needed for maturation and subsequent neurogenesis. These findings highlight the importance and complexity of reiterated Fgf signaling during cranial placode formation and subsequent sensory organ development.
Conflict of interest statement
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

