SIRT1/PARP1 crosstalk: connecting DNA damage and metabolism
- PMID: 24360018
- PMCID: PMC3898398
- DOI: 10.1186/2041-9414-4-6
SIRT1/PARP1 crosstalk: connecting DNA damage and metabolism
Abstract
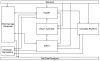
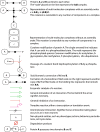
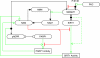
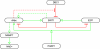
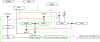
An intricate network regulates the activities of SIRT1 and PARP1 proteins and continues to be uncovered. Both SIRT1 and PARP1 share a common co-factor nicotinamide adenine dinucleotide (NAD+) and several common substrates, including regulators of DNA damage response and circadian rhythms. We review this complex network using an interactive Molecular Interaction Map (MIM) to explore the interplay between these two proteins. Here we discuss how NAD + competition and post-transcriptional/translational feedback mechanisms create a regulatory network sensitive to environmental cues, such as genotoxic stress and metabolic states, and examine the role of those interactions in DNA repair and ultimately, cell fate decisions.
Figures

Similar articles
-
Predicted Role of NAD Utilization in the Control of Circadian Rhythms during DNA Damage Response.PLoS Comput Biol. 2015 May 28;11(5):e1004144. doi: 10.1371/journal.pcbi.1004144. eCollection 2015 May. PLoS Comput Biol. 2015. PMID: 26020938 Free PMC article.
-
PBX1-SIRT1 Positive Feedback Loop Attenuates ROS-Mediated HF-MSC Senescence and Apoptosis.Stem Cell Rev Rep. 2023 Feb;19(2):443-454. doi: 10.1007/s12015-022-10425-w. Epub 2022 Aug 12. Stem Cell Rev Rep. 2023. PMID: 35962175 Free PMC article.
-
Poly(ADP-ribose) Polymerase (PARP) and PARP Inhibitors: Mechanisms of Action and Role in Cardiovascular Disorders.Cardiovasc Toxicol. 2018 Dec;18(6):493-506. doi: 10.1007/s12012-018-9462-2. Cardiovasc Toxicol. 2018. PMID: 29968072 Review.
-
Poly(ADP-ribose) polymerase 1 regulates mitochondrial DNA repair in an NAD-dependent manner.J Biol Chem. 2021 Jan-Jun;296:100309. doi: 10.1016/j.jbc.2021.100309. Epub 2021 Jan 19. J Biol Chem. 2021. PMID: 33482196 Free PMC article.
-
[Importance of NAMPT-mediated NAD-biosynthesis and NAD-dependent deacetylase SIRT1 in the crosstalk between circadian rhythm and metabolism].Nihon Rinsho. 2013 Dec;71(12):2187-93. Nihon Rinsho. 2013. PMID: 24437277 Review. Japanese.
Cited by
-
PARP1 Inhibition Augments UVB-Mediated Mitochondrial Changes-Implications for UV-Induced DNA Repair and Photocarcinogenesis.Cancers (Basel). 2019 Dec 18;12(1):5. doi: 10.3390/cancers12010005. Cancers (Basel). 2019. PMID: 31861350 Free PMC article.
-
Fueling genome maintenance: On the versatile roles of NAD+ in preserving DNA integrity.J Biol Chem. 2022 Jun;298(6):102037. doi: 10.1016/j.jbc.2022.102037. Epub 2022 May 17. J Biol Chem. 2022. PMID: 35595095 Free PMC article. Review.
-
Targeting the NAD+/SIRT1 axis: A metabolic strategy to overcome oxaliplatin resistance in colorectal cancer.World J Gastroenterol. 2025 Jun 7;31(21):106530. doi: 10.3748/wjg.v31.i21.106530. World J Gastroenterol. 2025. PMID: 40538510 Free PMC article.
-
HDACs and HDAC Inhibitors in Cancer Development and Therapy.Cold Spring Harb Perspect Med. 2016 Oct 3;6(10):a026831. doi: 10.1101/cshperspect.a026831. Cold Spring Harb Perspect Med. 2016. PMID: 27599530 Free PMC article. Review.
-
Relevance of SIRT1-NF-κB Axis as Therapeutic Target to Ameliorate Inflammation in Liver Disease.Int J Mol Sci. 2020 May 29;21(11):3858. doi: 10.3390/ijms21113858. Int J Mol Sci. 2020. PMID: 32485811 Free PMC article. Review.
References
-
- Blander G, Guarente L. The Sir2 family of protein deacetylases. Annu Rev Plant Physiol Plant Mol Biol. 2004;4:417–435. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
