SUMOylation inhibits FOXM1 activity and delays mitotic transition
- PMID: 24362530
- PMCID: PMC4096495
- DOI: 10.1038/onc.2013.546
SUMOylation inhibits FOXM1 activity and delays mitotic transition
Abstract
The forkhead box transcription factor FOXM1 is an essential effector of G2/M-phase transition, mitosis and the DNA damage response. As such, it is frequently deregulated during tumorigenesis. Here we report that FOXM1 is dynamically modified by SUMO1 but not by SUMO2/3 at multiple sites. We show that FOXM1 SUMOylation is enhanced in MCF-7 breast cancer cells in response to treatment with epirubicin and mitotic inhibitors. Mutation of five consensus conjugation motifs yielded a SUMOylation-deficient mutant FOXM1. Conversely, fusion of the E2 ligase Ubc9 to FOXM1 generated an auto-SUMOylating mutant (FOXM1-Ubc9). Analysis of wild-type FOXM1 and mutants revealed that SUMOylation inhibits FOXM1 activity, promotes translocation to the cytoplasm and enhances APC/Cdh1-mediated ubiquitination and degradation. Further, expression of the SUMOylation-deficient mutant enhanced cell proliferation compared with wild-type FOXM1, whereas the FOXM1-Ubc9 fusion protein resulted in persistent cyclin B1 expression and slowed the time from mitotic entry to exit. In summary, our findings suggest that SUMOylation attenuates FOXM1 activity and causes mitotic delay in cytotoxic drug response.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





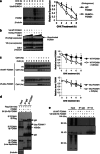


References
-
- Chien AJ, Moasser MM . Cellular mechanisms of resistance to anthracyclines and taxanes in cancer: intrinsic and acquired. Semin Oncol 2008; 35: S1–S14 quiz S39. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

