Pch2 is a hexameric ring ATPase that remodels the chromosome axis protein Hop1
- PMID: 24367111
- PMCID: PMC3890899
- DOI: 10.1073/pnas.1310755111
Pch2 is a hexameric ring ATPase that remodels the chromosome axis protein Hop1
Abstract
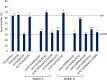
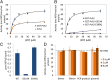
In budding yeast the pachytene checkpoint 2 (Pch2) protein regulates meiotic chromosome axis structure by maintaining the domain-like organization of the synaptonemal complex proteins homolog pairing 1 (Hop1) and molecular zipper 1 (Zip1). Pch2 has also been shown to modulate meiotic double-strand break repair outcomes to favor recombination between homologs, play an important role in the progression of meiotic recombination, and maintain ribosomal DNA stability. Pch2 homologs are present in fruit flies, worms, and mammals, however the molecular mechanism of Pch2 function is unknown. In this study we provide a unique and detailed biochemical analysis of Pch2. We find that purified Pch2 is an AAA+ (ATPases associated with diverse cellular activities) protein that oligomerizes into single hexameric rings in the presence of nucleotides. In addition, we show Pch2 binds to Hop1, a critical axial component of the synaptonemal complex that establishes interhomolog repair bias, in a nucleotide-dependent fashion. Importantly, we demonstrate that Pch2 displaces Hop1 from large DNA substrates and that both ATP binding and hydrolysis by Pch2 are required for Pch2-Hop1 transactions. Based on these and previous cell biological observations, we suggest that Pch2 impacts meiotic chromosome function by directly regulating Hop1 localization.
Keywords: AAA proteins; hexameric ATPase; meiosis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
Pch2(TRIP13): controlling cell division through regulation of HORMA domains.Chromosoma. 2015 Sep;124(3):333-9. doi: 10.1007/s00412-015-0516-y. Epub 2015 Apr 21. Chromosoma. 2015. PMID: 25895724 Review.
-
The Pch2 AAA+ ATPase promotes phosphorylation of the Hop1 meiotic checkpoint adaptor in response to synaptonemal complex defects.Nucleic Acids Res. 2016 Sep 19;44(16):7722-41. doi: 10.1093/nar/gkw506. Epub 2016 Jun 2. Nucleic Acids Res. 2016. PMID: 27257060 Free PMC article.
-
Pch2 acts through Xrs2 and Tel1/ATM to modulate interhomolog bias and checkpoint function during meiosis.PLoS Genet. 2011 Nov;7(11):e1002351. doi: 10.1371/journal.pgen.1002351. Epub 2011 Nov 3. PLoS Genet. 2011. PMID: 22072981 Free PMC article.
-
Homeostatic Control of Meiotic Prophase Checkpoint Function by Pch2 and Hop1.Curr Biol. 2020 Nov 16;30(22):4413-4424.e5. doi: 10.1016/j.cub.2020.08.064. Epub 2020 Sep 10. Curr Biol. 2020. PMID: 32916108
-
Getting there: understanding the chromosomal recruitment of the AAA+ ATPase Pch2/TRIP13 during meiosis.Curr Genet. 2021 Aug;67(4):553-565. doi: 10.1007/s00294-021-01166-3. Epub 2021 Mar 12. Curr Genet. 2021. PMID: 33712914 Free PMC article. Review.
Cited by
-
Mechanism and Control of Meiotic DNA Double-Strand Break Formation in S. cerevisiae.Front Cell Dev Biol. 2021 Mar 2;9:642737. doi: 10.3389/fcell.2021.642737. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 33748134 Free PMC article. Review.
-
Pch2(TRIP13): controlling cell division through regulation of HORMA domains.Chromosoma. 2015 Sep;124(3):333-9. doi: 10.1007/s00412-015-0516-y. Epub 2015 Apr 21. Chromosoma. 2015. PMID: 25895724 Review.
-
The multifaceted roles of the HORMA domain in cellular signaling.J Cell Biol. 2015 Nov 23;211(4):745-55. doi: 10.1083/jcb.201509076. J Cell Biol. 2015. PMID: 26598612 Free PMC article. Review.
-
The Arabidopsis Cdk1/Cdk2 homolog CDKA;1 controls chromosome axis assembly during plant meiosis.EMBO J. 2020 Feb 3;39(3):e101625. doi: 10.15252/embj.2019101625. Epub 2019 Sep 26. EMBO J. 2020. PMID: 31556459 Free PMC article.
-
Noncanonical Contributions of MutLγ to VDE-Initiated Crossovers During Saccharomyces cerevisiae Meiosis.G3 (Bethesda). 2019 May 7;9(5):1647-1654. doi: 10.1534/g3.119.400150. G3 (Bethesda). 2019. PMID: 30902890 Free PMC article.
References
-
- Hunter N (2007) Meiotic recombination. Molecular Genetics of Recombination: Topics in Current Genetics, eds Aguilera A, Rothstein R (Springer, Berlin), pp 381–442.
-
- Bishop DK, Zickler D. Early decision; meiotic crossover interference prior to stable strand exchange and synapsis. Cell. 2004;117(1):9–15. - PubMed
-
- Börner GV, Kleckner N, Hunter N. Crossover/noncrossover differentiation, synaptonemal complex formation, and regulatory surveillance at the leptotene/zygotene transition of meiosis. Cell. 2004;117(1):29–45. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

