Testis-specific Fank1 gene in knockdown mice produces oligospermia via apoptosis
- PMID: 24369145
- PMCID: PMC3901870
- DOI: 10.4103/1008-682X.122592
Testis-specific Fank1 gene in knockdown mice produces oligospermia via apoptosis
Abstract
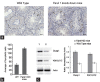
Fank1 is exclusively expressed in the testis from the meiosis phase to the haploid phase of spermatogenesis. In this study, we examined the function of Fank1 by establishing a Fank1-knockdown transgenic mouse model. The apoptotic statuses of the testes of the transgenic mice were tested using the terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) method. The FANK1 consensus DNA-binding sequence was identified using cyclic amplification of sequence target (CAST) analysis. Differentially expressed genes were examined using microarray analysis. A reduction in sperm number and an increase in apoptotic spermatocytes were observed in Fank1-knockdown mice, and the apoptotic cells were found to be primarily spermatogonia and spermatocytes. The CAST results demonstrated that the consensus DNA-binding sequence was AAAAAG, in which the percentage occurrence of each base at each position ranged from 55 to 86%. This sequence was present in the promoter regions of 10 differentially expressed genes that were examined using microarray analysis. In total, 17 genes were differentially expressed with changes in their expression levels greater than twofold. The abnormal expression of Fank1 target genes that were regulated directly or indirectly by Fank1 reduced the number of sperm in the knockdown mice. Thus, FANK1 may play a pivotal role in spermatogenesis as a transcription factor.
Figures



Similar articles
-
Normal spermatogenesis in Fank1 (fibronectin type 3 and ankyrin repeat domains 1) mutant mice.PeerJ. 2019 Apr 24;7:e6827. doi: 10.7717/peerj.6827. eCollection 2019. PeerJ. 2019. PMID: 31086747 Free PMC article.
-
Fank1 is a testis-specific gene encoding a nuclear protein exclusively expressed during the transition from the meiotic to the haploid phase of spermatogenesis.Gene Expr Patterns. 2007 Aug;7(7):777-83. doi: 10.1016/j.modgep.2007.05.005. Epub 2007 May 29. Gene Expr Patterns. 2007. PMID: 17604233
-
Fank1 interacts with Jab1 and regulates cell apoptosis via the AP-1 pathway.Cell Mol Life Sci. 2011 Jun;68(12):2129-39. doi: 10.1007/s00018-010-0559-4. Epub 2010 Oct 27. Cell Mol Life Sci. 2011. PMID: 20978819 Free PMC article.
-
RFX2 is a candidate downstream amplifier of A-MYB regulation in mouse spermatogenesis.BMC Dev Biol. 2009 Dec 9;9:63. doi: 10.1186/1471-213X-9-63. BMC Dev Biol. 2009. PMID: 20003220 Free PMC article.
-
The low expression of Dmrt7 is associated with spermatogenic arrest in cattle-yak.Mol Biol Rep. 2014 Nov;41(11):7255-63. doi: 10.1007/s11033-014-3611-x. Epub 2014 Jul 23. Mol Biol Rep. 2014. PMID: 25052188
Cited by
-
COP9 signalosome complex subunit 5, an IFT20 binding partner, is essential to maintain male germ cell survival and acrosome biogenesis†.Biol Reprod. 2020 Feb 12;102(1):233-247. doi: 10.1093/biolre/ioz154. Biol Reprod. 2020. PMID: 31373619 Free PMC article.
-
cba-miR-222-3p involved in photoperiod-induced apoptosis in testes of striped hamsters by targeting TRAF7.Integr Zool. 2025 Jul;20(4):836-849. doi: 10.1111/1749-4877.12918. Epub 2024 Oct 28. Integr Zool. 2025. PMID: 39466916 Free PMC article.
-
Candidate genes for infertility: an in-silico study based on cytogenetic analysis.BMC Med Genomics. 2022 Aug 2;15(1):170. doi: 10.1186/s12920-022-01320-x. BMC Med Genomics. 2022. PMID: 35918717 Free PMC article.
-
Normal spermatogenesis in Fank1 (fibronectin type 3 and ankyrin repeat domains 1) mutant mice.PeerJ. 2019 Apr 24;7:e6827. doi: 10.7717/peerj.6827. eCollection 2019. PeerJ. 2019. PMID: 31086747 Free PMC article.
-
The Ankyrin Repeat Domain 49 (ANKRD49) Augments Autophagy of Serum-Starved GC-1 Cells through the NF-κB Pathway.PLoS One. 2015 Jun 4;10(6):e0128551. doi: 10.1371/journal.pone.0128551. eCollection 2015. PLoS One. 2015. PMID: 26043108 Free PMC article.
References
-
- Eddy EM, O’Brien DA. Gene expression during mammalian meiosis. Curr Top Dev Biol. 1998;37:141–200. - PubMed
-
- Hecht NB. Molecular mechanisms of male germ cell differentiation. Bioessays. 1998;20:555–61. - PubMed
-
- Zheng Z, Zheng H, Yan W. Fank1 is a testis-specific gene encoding a nuclear protein exclusively expressed during the transition from the meiotic to the haploid phase of spermatogenesis. Gene Expr Patterns. 2007;7:777–83. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

