Characterization of protein unfolding with solid-state nanopores
- PMID: 24370259
- PMCID: PMC4188535
- DOI: 10.2174/09298665113209990077
Characterization of protein unfolding with solid-state nanopores
Abstract
In this work, we review the process of protein unfolding characterized by a solid-state nanopore based device. The occupied or excluded volume of a protein molecule in a nanopore depends on the protein's conformation or shape. A folded protein has a larger excluded volume in a nanopore thus it blocks more ionic current flow than its unfolded form and produces a greater current blockage amplitude. The time duration a protein stays in a pore also depends on the protein's folding state. We use Bovine Serum Albumin (BSA) as a model protein to discuss this current blockage amplitude and the time duration associated with the protein unfolding process. BSA molecules were measured in folded, partially unfolded, and completely unfolded conformations in solid-state nanopores. We discuss experimental results, data analysis, and theoretical considerations of BSA protein unfolding measured with silicon nitride nanopores. We show this nanopore method is capable of characterizing a protein's unfolding process at single molecule level. Problems and future studies in characterization of protein unfolding using a solid-state nanopore device will also be discussed.
Figures




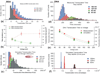
References
-
- Dobson CM. Protein folding and misfolding. Nature. 2003;426:884–890. - PubMed
-
- Daggett V, Levitt M. Protein Unfolding Pathways Explored through Molecular Dynamics Simulations. J. Mol. Biol. 1993;(232):600–619. - PubMed
-
- Oukhaled G, Mathe J, Biance A-L, Bacri L, Betton J-M, Lairez D, Pelta J, Auvray L. Unfolding of Proteins and Long Transient Conformations Detected by Single Nanopore Recording. Phys. Rev. Lett. 2007;98 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
