The ancient small GTPase Rab21 functions in intermediate endocytic steps in trypanosomes
- PMID: 24376004
- PMCID: PMC3910970
- DOI: 10.1128/EC.00269-13
The ancient small GTPase Rab21 functions in intermediate endocytic steps in trypanosomes
Abstract
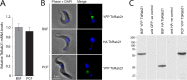
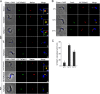
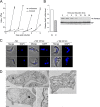
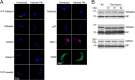
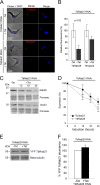
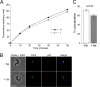
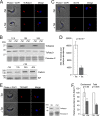
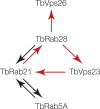
Endocytosis is an essential process in nearly all eukaryotic cells, including the African trypanosome Trypanosoma brucei. Endocytosis in these organisms is exclusively clathrin mediated, although several lineage-specific features indicate that precise mechanisms are distinct from those of higher eukaryotes. T. brucei Rab21 is a member of an ancient, pan-eukaryotic, endocytic Rab clade that is retained by trypanosomes. We show that T. brucei Rab21 (TbRab21) localizes to endosomes, partially colocalizing with TbRab5A, TbRab28, and TbVps23, the latter two being present at late endosomes. TbRab21 expression is essential for cellular proliferation, and its suppression results in a partial block in traffic to the lysosome. RNA interference (RNAi)-mediated knockdown of TbRab21 had no effect on TbRab5A expression or location but did result in decreased in trans expression of ESCRT (trypanosome endosomal sorting complex required for transport) components and TbRab28, while knockdown of ESCRT subunit TbVps23 resulted in decreased TbRab21 expression. These data suggest that TbRab21 acts downstream of TbRab5A and functions in intimate connection with the trypanosome ESCRT system.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

