Subtilase SprP exerts pleiotropic effects in Pseudomonas aeruginosa
- PMID: 24376018
- PMCID: PMC3937732
- DOI: 10.1002/mbo3.150
Subtilase SprP exerts pleiotropic effects in Pseudomonas aeruginosa
Abstract
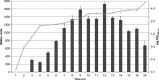
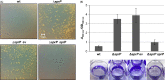
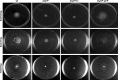
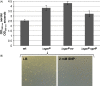
The open reading frame PA1242 in the genome of Pseudomonas aeruginosa PAO1 encodes a putative protease belonging to the peptidase S8 family of subtilases. The respective enzyme termed SprP consists of an N-terminal signal peptide and a so-called S8 domain linked by a domain of unknown function (DUF). Presumably, this DUF domain defines a discrete class of Pseudomonas proteins as homologous domains can be identified almost exclusively in proteins of the genus Pseudomonas. The sprP gene was expressed in Escherichia coli and proteolytic activity was demonstrated. A P. aeruginosa ∆sprP mutant was constructed and its gene expression pattern compared to the wild-type strain by genome microarray analysis revealing altered expression levels of 218 genes. Apparently, SprP is involved in regulation of a variety of different cellular processes in P. aeruginosa including pyoverdine synthesis, denitrification, the formation of cell aggregates, and of biofilms.
Keywords: Biofilm; Pseudomonas aeruginosa; Pyoverdine.; microarray; motility; orf PA1242; protease.
© 2013 The Authors. MicrobiologyOpen published by John Wiley & Sons Ltd.
Figures



Similar articles
-
Functional expression, purification, and biochemical properties of subtilase SprP from Pseudomonas aeruginosa.Microbiologyopen. 2015 Oct;4(5):743-52. doi: 10.1002/mbo3.275. Epub 2015 Jul 14. Microbiologyopen. 2015. PMID: 26175208 Free PMC article.
-
PA2663 (PpyR) increases biofilm formation in Pseudomonas aeruginosa PAO1 through the psl operon and stimulates virulence and quorum-sensing phenotypes.Appl Microbiol Biotechnol. 2008 Feb;78(2):293-307. doi: 10.1007/s00253-007-1308-y. Epub 2007 Dec 22. Appl Microbiol Biotechnol. 2008. PMID: 18157527
-
Cloning and sequence analysis of an EnvCD homologue in Pseudomonas aeruginosa: regulation by iron and possible involvement in the secretion of the siderophore pyoverdine.Mol Microbiol. 1993 Nov;10(3):529-44. doi: 10.1111/j.1365-2958.1993.tb00925.x. Mol Microbiol. 1993. PMID: 7968531
-
Membrane-association determinants of the omega-amino acid monooxygenase PvdA, a pyoverdine biosynthetic enzyme from Pseudomonas aeruginosa.Microbiology (Reading). 2008 Sep;154(Pt 9):2804-2813. doi: 10.1099/mic.0.2008/018804-0. Microbiology (Reading). 2008. PMID: 18757814
-
Cloning and nucleotide sequence of Pseudomonas aeruginosa DNA gyrase gyrA gene from strain PAO1 and quinolone-resistant clinical isolates.Antimicrob Agents Chemother. 1994 Sep;38(9):1944-52. doi: 10.1128/AAC.38.9.1944. Antimicrob Agents Chemother. 1994. PMID: 7811002 Free PMC article.
Cited by
-
Ethanol Stimulates Trehalose Production through a SpoT-DksA-AlgU-Dependent Pathway in Pseudomonas aeruginosa.J Bacteriol. 2019 May 22;201(12):e00794-18. doi: 10.1128/JB.00794-18. Print 2019 Jun 15. J Bacteriol. 2019. PMID: 30936375 Free PMC article.
-
Metagenomic discovery of novel enzymes and biosurfactants in a slaughterhouse biofilm microbial community.Sci Rep. 2016 Jun 8;6:27035. doi: 10.1038/srep27035. Sci Rep. 2016. PMID: 27271534 Free PMC article.
-
An optogenetic toolbox of LOV-based photosensitizers for light-driven killing of bacteria.Sci Rep. 2018 Oct 9;8(1):15021. doi: 10.1038/s41598-018-33291-4. Sci Rep. 2018. PMID: 30301917 Free PMC article.
-
Functional expression, purification, and biochemical properties of subtilase SprP from Pseudomonas aeruginosa.Microbiologyopen. 2015 Oct;4(5):743-52. doi: 10.1002/mbo3.275. Epub 2015 Jul 14. Microbiologyopen. 2015. PMID: 26175208 Free PMC article.
-
Outer membrane vesicles for vaccination and targeted drug delivery.Wiley Interdiscip Rev Nanomed Nanobiotechnol. 2019 Mar;11(2):e1523. doi: 10.1002/wnan.1523. Epub 2018 Apr 26. Wiley Interdiscip Rev Nanomed Nanobiotechnol. 2019. PMID: 29701017 Free PMC article. Review.
References
-
- Alexeyev MF, Shokolenko IN, Croughan TP. Improved antibiotic-resistance gene cassettes and omega elements for Escherichia coli vector construction and in vitro deletion/insertion mutagenesis. Gene. 1995;160:63–67. - PubMed
-
- Antao CM, Malcata FX. Plant serine proteases: biochemical, physiological and molecular features. Plant Physiol. Biochem. 2005;43:637–650. - PubMed
-
- Benjamini Y, Hochberg Y. Controlling the fasle discovery rate – a practical and powerful approach to multiple testing. J. Roy. Stat. Soc. 1995;57:289–300.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

