Circular Permutation of the Trp-cage: Fold Rescue upon Addition of a Hydrophobic Staple
- PMID: 24376912
- PMCID: PMC3870897
- DOI: 10.1039/C3RA43674H
Circular Permutation of the Trp-cage: Fold Rescue upon Addition of a Hydrophobic Staple
Abstract
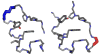
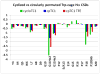
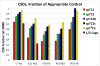
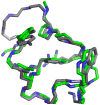
The Trp-cage, at 20 residues in length, is generally acknowledged as the smallest fully protein-like folding motif. Linking the termini by a two-residue unit and excising one residue affords circularly permuted sequences that adopt the same structure. This represents the first successful circular permutation of any fold of less than 50-residue length. As was observed for the original topology, a hydrophobic staple near the chain termini is required for enhanced fold stability.
Figures
References
-
- Lo WC, Lee CC, Lee CY, Lyu PC. Nucleic Acids Res. 2009;37:D328. CPDB (Circular Permutation Data Base) is found under http://sarst.life.nthu.edu.tw/cpdb/ - PMC - PubMed
-
- Kier B, Byrne A, Scian M, Anderson J, Andersen NH. Folding landscape exploration by circular permutation and capping β structures. In: Kokotos G, Constantinou V, Matsoukas J, editors. Peptides 2012, Proceeding of the 32ndEPS. 2012. pp. 70–71.
-
- Neidigh JW, Fesinmeyer RM, Andersen NH. Nat Struct Biol. 2002;9:425. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
