Discovery and saturation analysis of cancer genes across 21 tumour types
- PMID: 24390350
- PMCID: PMC4048962
- DOI: 10.1038/nature12912
Discovery and saturation analysis of cancer genes across 21 tumour types
Abstract
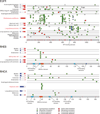
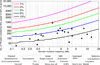
Although a few cancer genes are mutated in a high proportion of tumours of a given type (>20%), most are mutated at intermediate frequencies (2-20%). To explore the feasibility of creating a comprehensive catalogue of cancer genes, we analysed somatic point mutations in exome sequences from 4,742 human cancers and their matched normal-tissue samples across 21 cancer types. We found that large-scale genomic analysis can identify nearly all known cancer genes in these tumour types. Our analysis also identified 33 genes that were not previously known to be significantly mutated in cancer, including genes related to proliferation, apoptosis, genome stability, chromatin regulation, immune evasion, RNA processing and protein homeostasis. Down-sampling analysis indicates that larger sample sizes will reveal many more genes mutated at clinically important frequencies. We estimate that near-saturation may be achieved with 600-5,000 samples per tumour type, depending on background mutation frequency. The results may help to guide the next stage of cancer genomics.
Figures



References
-
- Garraway LA, Lander ES. Lessons from the cancer genome. Cell. 2013;153:17–37. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

