A multi-scale model of hepcidin promoter regulation reveals factors controlling systemic iron homeostasis
- PMID: 24391488
- PMCID: PMC3879105
- DOI: 10.1371/journal.pcbi.1003421
A multi-scale model of hepcidin promoter regulation reveals factors controlling systemic iron homeostasis
Abstract
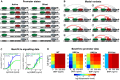
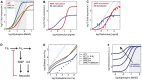
Systemic iron homeostasis involves a negative feedback circuit in which the expression level of the peptide hormone hepcidin depends on and controls the iron blood levels. Hepcidin expression is regulated by the BMP6/SMAD and IL6/STAT signaling cascades. Deregulation of either pathway causes iron-related diseases such as hemochromatosis or anemia of inflammation. We quantitatively analyzed how BMP6 and IL6 control hepcidin expression. Transcription factor (TF) phosphorylation and reporter gene expression were measured under co-stimulation conditions, and the promoter was perturbed by mutagenesis. Using mathematical modeling, we systematically analyzed potential mechanisms of cooperative and competitive promoter regulation by the transcription factors, and experimentally validated the model predictions. Our results reveal that hepcidin cross-regulation primarily occurs by combinatorial transcription factor binding to the promoter, whereas signaling crosstalk is insignificant. We find that the presence of two BMP-responsive elements enhances the steepness of the promoter response towards the iron-sensing BMP signaling axis, which promotes iron homeostasis in vivo. IL6 co-stimulation reduces the promoter sensitivity towards the BMP signal, because the SMAD and STAT transcription factors compete for recruiting RNA polymerase to the transcription start site. This may explain why inflammatory signals disturb iron homeostasis in anemia of inflammation. Taken together, our results reveal why the iron homeostasis circuit is sensitive to perturbations implicated in disease.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Bone morphogenetic protein (BMP)-responsive elements located in the proximal and distal hepcidin promoter are critical for its response to HJV/BMP/SMAD.J Mol Med (Berl). 2009 May;87(5):471-80. doi: 10.1007/s00109-009-0447-2. Epub 2009 Feb 20. J Mol Med (Berl). 2009. PMID: 19229506
-
Two BMP responsive elements, STAT, and bZIP/HNF4/COUP motifs of the hepcidin promoter are critical for BMP, SMAD1, and HJV responsiveness.Blood. 2009 Jan 15;113(3):688-95. doi: 10.1182/blood-2008-05-160184. Epub 2008 Nov 7. Blood. 2009. PMID: 18997172 Free PMC article.
-
Hepcidin and the BMP-SMAD pathway: An unexpected liaison.Vitam Horm. 2019;110:71-99. doi: 10.1016/bs.vh.2019.01.004. Epub 2019 Feb 10. Vitam Horm. 2019. PMID: 30798817 Review.
-
Hepatic heparan sulfate is a master regulator of hepcidin expression and iron homeostasis in human hepatocytes and mice.J Biol Chem. 2019 Sep 6;294(36):13292-13303. doi: 10.1074/jbc.RA118.007213. Epub 2019 Jul 17. J Biol Chem. 2019. PMID: 31315930 Free PMC article.
-
Modulation of hepcidin to treat iron deregulation: potential clinical applications.Expert Rev Hematol. 2016;9(2):169-86. doi: 10.1586/17474086.2016.1124757. Epub 2015 Dec 15. Expert Rev Hematol. 2016. PMID: 26669208 Free PMC article. Review.
Cited by
-
Pharmacological Targeting of BMP6-SMAD Mediated Hepcidin Expression Does Not Improve the Outcome of Systemic Infections With Intra-Or Extracellular Gram-Negative Bacteria in Mice.Front Cell Infect Microbiol. 2021 Jul 23;11:705087. doi: 10.3389/fcimb.2021.705087. eCollection 2021. Front Cell Infect Microbiol. 2021. PMID: 34368018 Free PMC article.
-
Novel loci affecting iron homeostasis and their effects in individuals at risk for hemochromatosis.Nat Commun. 2014 Oct 29;5:4926. doi: 10.1038/ncomms5926. Nat Commun. 2014. PMID: 25352340 Free PMC article.
-
Signal integration by the CYP1A1 promoter--a quantitative study.Nucleic Acids Res. 2015 Jun 23;43(11):5318-30. doi: 10.1093/nar/gkv423. Epub 2015 May 1. Nucleic Acids Res. 2015. PMID: 25934798 Free PMC article.
-
Exon Definition Facilitates Reliable Control of Alternative Splicing in the RON Proto-Oncogene.Biophys J. 2020 Apr 21;118(8):2027-2041. doi: 10.1016/j.bpj.2020.02.022. Epub 2020 Mar 3. Biophys J. 2020. PMID: 32336349 Free PMC article.
-
Modelling Systemic Iron Regulation during Dietary Iron Overload and Acute Inflammation: Role of Hepcidin-Independent Mechanisms.PLoS Comput Biol. 2017 Jan 9;13(1):e1005322. doi: 10.1371/journal.pcbi.1005322. eCollection 2017 Jan. PLoS Comput Biol. 2017. PMID: 28068331 Free PMC article.
References
-
- Hentze MW, Muckenthaler MU, Galy B, Camaschella C (2010) Two to tango: regulation of Mammalian iron metabolism. Cell 142: 24–38 - PubMed
-
- Becskei A, Serrano L (2000) Engineering stability in gene networks by autoregulation. Nature 405: 590–593 - PubMed
-
- Paulsen M, Legewie S, Eils R, Karaulanov E, Niehrs C (2011) Negative feedback in the bone morphogenetic protein 4 (BMP4) synexpression group governs its dynamic signaling range and canalizes development. Proceedings of the National Academy of Sciences of the United States of America 108: 10202–10207 - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

