Myogenic differentiation of muscular dystrophy-specific induced pluripotent stem cells for use in drug discovery
- PMID: 24396035
- PMCID: PMC3925053
- DOI: 10.5966/sctm.2013-0095
Myogenic differentiation of muscular dystrophy-specific induced pluripotent stem cells for use in drug discovery
Abstract
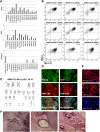
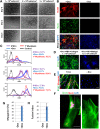
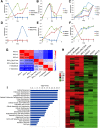
Human induced pluripotent stem cells (iPSCs) represent a scalable source of potentially any cell type for disease modeling and therapeutic screening. We have a particular interest in modeling skeletal muscle from various genetic backgrounds; however, efficient and reproducible methods for the myogenic differentiation of iPSCs have not previously been demonstrated. Ectopic myogenic differentiation 1 (MyoD) expression has been shown to induce myogenesis in primary cell types, but the same effect has been unexpectedly challenging to reproduce in human iPSCs. In this study, we report that optimization of culture conditions enabled direct MyoD-mediated differentiation of iPSCs into myoblasts without the need for an intermediate step or cell sorting. MyoD induction mediated efficient cell fusion of mature myocytes yielding multinucleated myosin heavy chain-positive myotubes. We applied the same approach to dystrophic iPSCs, generating 16 iPSC lines from fibroblasts of four patients with Duchenne and Becker muscular dystrophies. As seen with iPSCs from healthy donors, within 36 hours from MyoD induction there was a clear commitment toward the myogenic identity by the majority of iPSCs in culture (50%-70%). The patient iPSC-derived myotubes successfully adopted the skeletal muscle program, as determined by global gene expression profiling, and were functionally responsive to treatment with hypertrophic proteins insulin-like growth factor 1 (IGF-1) and wingless-type MMTV integration site family, member 7A (Wnt7a), which are being investigated as potential treatments for muscular dystrophy in clinical and preclinical studies, respectively. Our results demonstrate that iPSCs have no intrinsic barriers preventing MyoD from inducing efficient and rapid myogenesis and thus providing a scalable source of normal and dystrophic myoblasts for use in disease modeling and drug discovery.
Keywords: Differentiation; Induced pluripotent stem cells; Muscular dystrophy; Skeletal muscle.
Figures


Comment in
-
Regeneration: making muscle from hPSCs.Cell Res. 2014 Oct;24(10):1159-61. doi: 10.1038/cr.2014.91. Epub 2014 Jul 8. Cell Res. 2014. PMID: 25001388 Free PMC article.
References
-
- Heslop L, Morgan JE, Partridge TA. Evidence for a myogenic stem cell that is exhausted in dystrophic muscle. J Cell Sci. 2000;113:2299–2308. - PubMed
-
- Yablonka-Reuveni Z, Anderson JE. Satellite cells from dystrophic (mdx) mice display accelerated differentiation in primary cultures and in isolated myofibers. Dev Dyn. 2006;235:203–212. - PubMed
-
- Barberi T, Bradbury M, Dincer Z, et al. Derivation of engraftable skeletal myoblasts from human embryonic stem cells. Nat Med. 2007;13:642–648. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

