Myosin binding protein-C slow: a multifaceted family of proteins with a complex expression profile in fast and slow twitch skeletal muscles
- PMID: 24399972
- PMCID: PMC3872291
- DOI: 10.3389/fphys.2013.00391
Myosin binding protein-C slow: a multifaceted family of proteins with a complex expression profile in fast and slow twitch skeletal muscles
Abstract
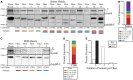
Myosin Binding Protein-C slow (sMyBP-C) comprises a complex family of proteins expressed in slow and fast type skeletal muscles. Similar to its fast and cardiac counterparts, sMyBP-C functions to modulate the formation of actomyosin cross-bridges, and to organize and stabilize sarcomeric A- and M-bands. The slow form of MyBP-C was originally classified as a single protein, however several variants encoded by the single MYBPC1 gene have been recently identified. Alternative splicing of the 5' and 3' ends of the MYBPC1 transcript has led to the differential expression of small unique segments interspersed between common domains. In addition, the NH2-terminus of sMyBP-C undergoes complex phosphorylation. Thus, alternative splicing and phosphorylation appear to regulate the functional activities of sMyBP-C. sMyBP-C proteins are not restricted to slow twitch muscles, but they are abundantly expressed in fast twitch muscles, too. Using bioinformatic tools, we herein perform a systematic comparison of the known human and mouse sMyBP-C variants. In addition, using single fiber westerns and antibodies to a common region of all known sMyBP-C variants, we present a detailed and comprehensive characterization of the expression profile of sMyBP-C proteins in the slow twitch soleus and the fast twitch flexor digitorum brevis (FDB) mouse muscles. Our studies demonstrate for the first time that distinct sMyBP-C variants are co-expressed in the same fiber, and that their expression profile differs among fibers. Given the differential expression of sMyBP-C variants in single fibers, it becomes apparent that each variant or combination thereof may play unique roles in the regulation of actomyosin cross-bridges formation and the stabilization of thick filaments.
Keywords: MYBPC1; single fiber expression; skeletal muscle.
Figures



References
-
- Ackermann M. A., Hu L.-Y. R., Bowman A. L., Bloch R. J., Kontrogianni-Konstantopoulos A. (2009). Obscurin interacts with a novel isoform of MyBP-C slow at the periphery of the sarcomeric M-band and regulates thick filament assembly. Mol. Biol. Cell 20, 2963–2978 10.1091/mbc.E08-12-1251 - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

