A conserved role for Snail as a potentiator of active transcription
- PMID: 24402316
- PMCID: PMC3909790
- DOI: 10.1101/gad.230953.113
A conserved role for Snail as a potentiator of active transcription
Abstract
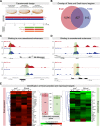
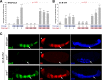
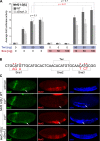
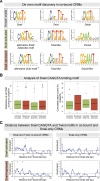
The transcription factors of the Snail family are key regulators of epithelial-mesenchymal transitions, cell morphogenesis, and tumor metastasis. Since its discovery in Drosophila ∼25 years ago, Snail has been extensively studied for its role as a transcriptional repressor. Here we demonstrate that Drosophila Snail can positively modulate transcriptional activation. By combining information on in vivo occupancy with expression profiling of hand-selected, staged snail mutant embryos, we identified 106 genes that are potentially directly regulated by Snail during mesoderm development. In addition to the expected Snail-repressed genes, almost 50% of Snail targets showed an unanticipated activation. The majority of "Snail-activated" genes have enhancer elements cobound by Twist and are expressed in the mesoderm at the stages of Snail occupancy. Snail can potentiate Twist-mediated enhancer activation in vitro and is essential for enhancer activity in vivo. Using a machine learning approach, we show that differentially enriched motifs are sufficient to predict Snail's regulatory response. In silico mutagenesis revealed a likely causative motif, which we demonstrate is essential for enhancer activation. Taken together, these data indicate that Snail can potentiate enhancer activation by collaborating with different activators, providing a new mechanism by which Snail regulates development.
Keywords: Drosophila embryogenesis; Snail; Twist; activation; repression; spatiotemporal gene expression; transcription factor.
Figures

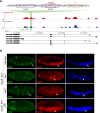
Similar articles
-
The myogenic repressor gene Holes in muscles is a direct transcriptional target of Twist and Tinman in the Drosophila embryonic mesoderm.Dev Biol. 2015 Apr 15;400(2):266-76. doi: 10.1016/j.ydbio.2015.02.005. Epub 2015 Feb 20. Dev Biol. 2015. PMID: 25704510 Free PMC article.
-
High conservation of transcription factor binding and evidence for combinatorial regulation across six Drosophila species.Nat Genet. 2011 May;43(5):414-20. doi: 10.1038/ng.808. Epub 2011 Apr 10. Nat Genet. 2011. PMID: 21478888
-
Identification of novel genes in Drosophila reveals the complex regulation of early gene activity in the mesoderm.Proc Natl Acad Sci U S A. 1996 Sep 17;93(19):10327-32. doi: 10.1073/pnas.93.19.10327. Proc Natl Acad Sci U S A. 1996. PMID: 8816799 Free PMC article.
-
Is mechano-sensitive expression of twist involved In mesoderm formation?Biol Cell. 2004 Sep;96(7):471-7. doi: 10.1016/j.biolcel.2004.04.009. Biol Cell. 2004. PMID: 15380614 Review.
-
Snail transcription factors in hematopoietic cell development: a model of functional redundancy.Exp Hematol. 2014 Jun;42(6):425-30. doi: 10.1016/j.exphem.2014.03.002. Epub 2014 Mar 24. Exp Hematol. 2014. PMID: 24674754 Free PMC article. Review.
Cited by
-
A coherent FOXO3-SNAI2 feed-forward loop in autophagy.Proc Natl Acad Sci U S A. 2022 Mar 15;119(11):e2118285119. doi: 10.1073/pnas.2118285119. Epub 2022 Mar 10. Proc Natl Acad Sci U S A. 2022. PMID: 35271390 Free PMC article.
-
Context-dependent EMT programs in cancer metastasis.J Exp Med. 2019 May 6;216(5):1016-1026. doi: 10.1084/jem.20181827. Epub 2019 Apr 11. J Exp Med. 2019. PMID: 30975895 Free PMC article. Review.
-
Modulation of protein-DNA binding reveals mechanisms of spatiotemporal gene control in early Drosophila embryos.Elife. 2023 Nov 7;12:e85997. doi: 10.7554/eLife.85997. Elife. 2023. PMID: 37934571 Free PMC article.
-
Myosin-dependent remodeling of adherens junctions protects junctions from Snail-dependent disassembly.J Cell Biol. 2016 Jan 18;212(2):219-29. doi: 10.1083/jcb.201508056. Epub 2016 Jan 11. J Cell Biol. 2016. PMID: 26754645 Free PMC article.
-
Mammary Stem Cell Self-Renewal Is Regulated by Slit2/Robo1 Signaling through SNAI1 and mINSC.Cell Rep. 2015 Oct 13;13(2):290-301. doi: 10.1016/j.celrep.2015.09.006. Epub 2015 Oct 1. Cell Rep. 2015. PMID: 26440891 Free PMC article.
References
-
- Ashburner M, Thompson P, Roote J, Lasko PF, Grau Y, el Messal M, Roth S, Simpson P 1990. The genetics of a small autosomal region of Drosophila melanogaster containing the structural gene for alcohol dehydrogenase. VII. Characterization of the region around the snail and cactus loci. Genetics 126: 679–694 - PMC - PubMed
-
- Barrallo-Gimeno A, Nieto MA 2005. The Snail genes as inducers of cell movement and survival: Implications in development and cancer. Development 132: 3151–3161 - PubMed
-
- Bodmer R, Jan LY, Jan YN 1990. A new homeobox-containing gene, msh-2, is transiently expressed early during mesoderm formation of Drosophila. Development 110: 661–669 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
