Computationally designed libraries for rapid enzyme stabilization
- PMID: 24402331
- PMCID: PMC3893934
- DOI: 10.1093/protein/gzt061
Computationally designed libraries for rapid enzyme stabilization
Abstract
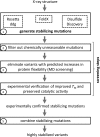
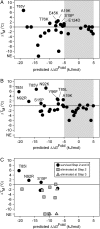
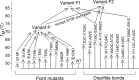
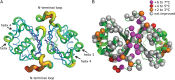
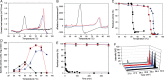
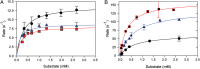
The ability to engineer enzymes and other proteins to any desired stability would have wide-ranging applications. Here, we demonstrate that computational design of a library with chemically diverse stabilizing mutations allows the engineering of drastically stabilized and fully functional variants of the mesostable enzyme limonene epoxide hydrolase. First, point mutations were selected if they significantly improved the predicted free energy of protein folding. Disulfide bonds were designed using sampling of backbone conformational space, which tripled the number of experimentally stabilizing disulfide bridges. Next, orthogonal in silico screening steps were used to remove chemically unreasonable mutations and mutations that are predicted to increase protein flexibility. The resulting library of 64 variants was experimentally screened, which revealed 21 (pairs of) stabilizing mutations located both in relatively rigid and in flexible areas of the enzyme. Finally, combining 10-12 of these confirmed mutations resulted in multi-site mutants with an increase in apparent melting temperature from 50 to 85°C, enhanced catalytic activity, preserved regioselectivity and a >250-fold longer half-life. The developed Framework for Rapid Enzyme Stabilization by Computational libraries (FRESCO) requires far less screening than conventional directed evolution.
Keywords: enzyme stability; in silico design; in silico screening; protein stability engineering; thermostability.
Figures

Similar articles
-
Stabilization of cyclohexanone monooxygenase by computational and experimental library design.Biotechnol Bioeng. 2019 Sep;116(9):2167-2177. doi: 10.1002/bit.27022. Epub 2019 Jun 24. Biotechnol Bioeng. 2019. PMID: 31124128 Free PMC article.
-
QM/MM study of the mechanism of enzymatic limonene 1,2-epoxide hydrolysis.Biochim Biophys Acta. 2012 Feb;1824(2):263-8. doi: 10.1016/j.bbapap.2011.08.014. Epub 2011 Sep 3. Biochim Biophys Acta. 2012. PMID: 21925621
-
X-ray crystallographic validation of structure predictions used in computational design for protein stabilization.Proteins. 2015 May;83(5):940-51. doi: 10.1002/prot.24791. Epub 2015 Mar 25. Proteins. 2015. PMID: 25739581
-
Structure of Rhodococcus erythropolis limonene-1,2-epoxide hydrolase reveals a novel active site.EMBO J. 2003 Jun 2;22(11):2583-92. doi: 10.1093/emboj/cdg275. EMBO J. 2003. PMID: 12773375 Free PMC article.
-
Prediction and analysis of structure, stability and unfolding of thermolysin-like proteases.J Comput Aided Mol Des. 1993 Aug;7(4):367-96. doi: 10.1007/BF02337558. J Comput Aided Mol Des. 1993. PMID: 8229092 Review.
Cited by
-
Stabilizing AqdC, a Pseudomonas Quinolone Signal-Cleaving Dioxygenase from Mycobacteria, by FRESCO-Based Protein Engineering.Chembiochem. 2021 Feb 15;22(4):733-742. doi: 10.1002/cbic.202000641. Epub 2020 Nov 16. Chembiochem. 2021. PMID: 33058333 Free PMC article.
-
Approaching boiling point stability of an alcohol dehydrogenase through computationally-guided enzyme engineering.Elife. 2020 Mar 31;9:e54639. doi: 10.7554/eLife.54639. Elife. 2020. PMID: 32228861 Free PMC article.
-
Enzyme stabilization via computationally guided protein stapling.Proc Natl Acad Sci U S A. 2017 Nov 21;114(47):12472-12477. doi: 10.1073/pnas.1708907114. Epub 2017 Nov 6. Proc Natl Acad Sci U S A. 2017. PMID: 29109284 Free PMC article.
-
Computational tools for the evaluation of laboratory-engineered biocatalysts.Chem Commun (Camb). 2016 Dec 22;53(2):284-297. doi: 10.1039/c6cc06055b. Chem Commun (Camb). 2016. PMID: 27812570 Free PMC article. Review.
-
Exploring Solanum tuberosum Epoxide Hydrolase Internal Architecture by Water Molecules Tracking.Biomolecules. 2018 Nov 12;8(4):143. doi: 10.3390/biom8040143. Biomolecules. 2018. PMID: 30424576 Free PMC article.
References
-
- Berendsen H.J.C., Postma J.P.M., Van Gunsteren W.F., Dinola A., Haak J.R. J. Chem. Phys. 1984;81:3684–3690.
-
- Besenmatter W., Kast P., Hilvert D. Protein Struct. Funct. Bioinform. 2007;66:500–506. - PubMed
-
- Blair M., Andrews P.C., Fraser B.H., Forsyth C.M., Junk P.C., Massi M., Tuck K.L. Synthesis. 2007;2007:1523–1527.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

