Transcriptome in vivo analysis (TIVA) of spatially defined single cells in live tissue
- PMID: 24412976
- PMCID: PMC3964595
- DOI: 10.1038/nmeth.2804
Transcriptome in vivo analysis (TIVA) of spatially defined single cells in live tissue
Abstract
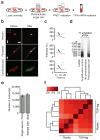
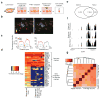
Transcriptome profiling of single cells resident in their natural microenvironment depends upon RNA capture methods that are both noninvasive and spatially precise. We engineered a transcriptome in vivo analysis (TIVA) tag, which upon photoactivation enables mRNA capture from single cells in live tissue. Using the TIVA tag in combination with RNA sequencing (RNA-seq), we analyzed transcriptome variance among single neurons in culture and in mouse and human tissue in vivo. Our data showed that the tissue microenvironment shapes the transcriptomic landscape of individual cells. The TIVA methodology is, to our knowledge, the first noninvasive approach for capturing mRNA from live single cells in their natural microenvironment.
Figures


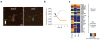
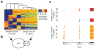
Comment in
-
Seeing is believing: new methods for in situ single-cell transcriptomics.Genome Biol. 2014 Mar 31;15(3):110. doi: 10.1186/gb4169. Genome Biol. 2014. PMID: 25000927 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

