Pathway-GPS and SIGORA: identifying relevant pathways based on the over-representation of their gene-pair signatures
- PMID: 24432194
- PMCID: PMC3883547
- DOI: 10.7717/peerj.229
Pathway-GPS and SIGORA: identifying relevant pathways based on the over-representation of their gene-pair signatures
Abstract
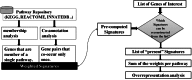
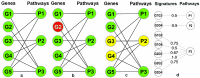
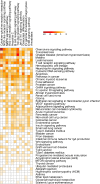
Motivation. Predominant pathway analysis approaches treat pathways as collections of individual genes and consider all pathway members as equally informative. As a result, at times spurious and misleading pathways are inappropriately identified as statistically significant, solely due to components that they share with the more relevant pathways. Results. We introduce the concept of Pathway Gene-Pair Signatures (Pathway-GPS) as pairs of genes that, as a combination, are specific to a single pathway. We devised and implemented a novel approach to pathway analysis, Signature Over-representation Analysis (SIGORA), which focuses on the statistically significant enrichment of Pathway-GPS in a user-specified gene list of interest. In a comparative evaluation of several published datasets, SIGORA outperformed traditional methods by delivering biologically more plausible and relevant results. Availability. An efficient implementation of SIGORA, as an R package with precompiled GPS data for several human and mouse pathway repositories is available for download from http://sigora.googlecode.com/svn/.
Keywords: Functional analysis; High-throughput data; Over-representation analysis; Pathway analysis; Shared components; Systems biology.
Figures




Similar articles
-
ADAGE signature analysis: differential expression analysis with data-defined gene sets.BMC Bioinformatics. 2017 Nov 22;18(1):512. doi: 10.1186/s12859-017-1905-4. BMC Bioinformatics. 2017. PMID: 29166858 Free PMC article.
-
Extending pathways based on gene lists using InterPro domain signatures.BMC Bioinformatics. 2008 Jan 4;9:3. doi: 10.1186/1471-2105-9-3. BMC Bioinformatics. 2008. PMID: 18177498 Free PMC article.
-
FELLA: an R package to enrich metabolomics data.BMC Bioinformatics. 2018 Dec 22;19(1):538. doi: 10.1186/s12859-018-2487-5. BMC Bioinformatics. 2018. PMID: 30577788 Free PMC article.
-
Retrospective analysis: reproducibility of interblastomere differences of mRNA expression in 2-cell stage mouse embryos is remarkably poor due to combinatorial mechanisms of blastomere diversification.Mol Hum Reprod. 2018 Jul 1;24(7):388-400. doi: 10.1093/molehr/gay021. Mol Hum Reprod. 2018. PMID: 29746690
-
Learning dysregulated pathways in cancers from differential variability analysis.Cancer Inform. 2014 Oct 23;13(Suppl 5):61-7. doi: 10.4137/CIN.S14066. eCollection 2014. Cancer Inform. 2014. PMID: 25392694 Free PMC article. Review.
Cited by
-
Systems Biology Approaches to Understanding the Human Immune System.Front Immunol. 2020 Jul 30;11:1683. doi: 10.3389/fimmu.2020.01683. eCollection 2020. Front Immunol. 2020. PMID: 32849587 Free PMC article.
-
Post-COVID symptoms are associated with endotypes reflecting poor inflammatory and hemostatic modulation.Front Immunol. 2023 Aug 23;14:1243689. doi: 10.3389/fimmu.2023.1243689. eCollection 2023. Front Immunol. 2023. PMID: 37680625 Free PMC article.
-
Development of a Blood-based Transcriptional Risk Score for Chronic Obstructive Pulmonary Disease.Am J Respir Crit Care Med. 2022 Jan 15;205(2):161-170. doi: 10.1164/rccm.202107-1584OC. Am J Respir Crit Care Med. 2022. PMID: 34739356 Free PMC article.
-
Comprehensive analysis of copy number aberrations in microsatellite stable colon cancer in view of stromal component.Br J Cancer. 2017 Jul 25;117(3):421-431. doi: 10.1038/bjc.2017.208. Epub 2017 Jul 6. Br J Cancer. 2017. PMID: 28683472 Free PMC article.
-
Comparative 'omics analyses differentiate Mycobacterium tuberculosis and Mycobacterium bovis and reveal distinct macrophage responses to infection with the human and bovine tubercle bacilli.Microb Genom. 2018 Mar;4(3):e000163. doi: 10.1099/mgen.0.000163. Epub 2018 Mar 20. Microb Genom. 2018. PMID: 29557774 Free PMC article.
References
-
- Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature Genetics. 2000;25(1):25–29. doi: 10.1038/75556. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

