Characterization of the water defect at the HIV-1 gp41 membrane spanning domain in bilayers with and without cholesterol using molecular simulations
- PMID: 24440660
- PMCID: PMC4040535
- DOI: 10.1016/j.bbamem.2014.01.009
Characterization of the water defect at the HIV-1 gp41 membrane spanning domain in bilayers with and without cholesterol using molecular simulations
Abstract
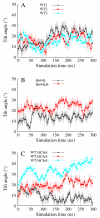
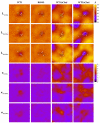
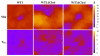
The membrane spanning domain (MSD) of human immunodeficiency virus 1 (HIV-1) envelope glycoprotein gp41 is important for fusion and infection. We used molecular dynamics (MD) simulations (3.4 μs total) to relate membrane and peptide properties that lead to water solvation of the α-helical gp41 MSD's midspan arginine in pure dipalmitoylphosphatidylcholine (DPPC) and in 50/50 DPPC/cholesterol membranes. We find that the midspan arginine is solvated by water that penetrates the inner leaflet, leading to a so-called water defect. The water defect is surprisingly robust across initial conditions and membrane compositions, but the presence of cholesterol modulates its behavior in several key ways. In the cholesterol-containing membranes, fluctuations in membrane thickness and water penetration depth are localized near the midspan arginine, and the MSD helices display a tightly regulated tilt angle. In the cholesterol-free membranes, thickness fluctuations are not as strongly correlated to the peptide position and tilt angles vary significantly depending on protein position relative to boundaries between domains of differing thickness. Cholesterol in an HIV-1 viral membrane is required for infection. Therefore, this work suggests that the colocalized water defect and membrane thickness fluctuations in cholesterol-containing viral membranes play an important role in fusion by bringing the membrane closer to a stability limit that must be crossed for fusion to occur.
Keywords: Membrane simulation; Metadynamics; Midspan arginine; Molecular dynamics; Snorkeling; Transmembrane.
Copyright © 2014 Elsevier B.V. All rights reserved.
Figures

Similar articles
-
All-atom models of the membrane-spanning domain of HIV-1 gp41 from metadynamics.Biophys J. 2010 Nov 17;99(10):3438-44. doi: 10.1016/j.bpj.2010.09.054. Biophys J. 2010. PMID: 21081093 Free PMC article.
-
Cholesterol-Mediated Clustering of the HIV Fusion Protein gp41 in Lipid Bilayers.J Mol Biol. 2022 Jan 30;434(2):167345. doi: 10.1016/j.jmb.2021.167345. Epub 2021 Nov 8. J Mol Biol. 2022. PMID: 34762895 Free PMC article.
-
Cholesterol mediates membrane curvature during fusion events.Phys Rev Lett. 2012 Jun 8;108(23):238103. doi: 10.1103/PhysRevLett.108.238103. Epub 2012 Jun 5. Phys Rev Lett. 2012. PMID: 23003994 Free PMC article.
-
The influence of cholesterol on membrane protein structure, function, and dynamics studied by molecular dynamics simulations.Biochim Biophys Acta. 2015 Sep;1848(9):1783-95. doi: 10.1016/j.bbamem.2015.03.029. Epub 2015 Apr 1. Biochim Biophys Acta. 2015. PMID: 25839353 Review.
-
Arginine in membranes: the connection between molecular dynamics simulations and translocon-mediated insertion experiments.J Membr Biol. 2011 Jan;239(1-2):35-48. doi: 10.1007/s00232-010-9330-x. Epub 2010 Dec 3. J Membr Biol. 2011. PMID: 21127848 Free PMC article. Review.
Cited by
-
HIV-1 Env gp41 Transmembrane Domain Dynamics Are Modulated by Lipid, Water, and Ion Interactions.Biophys J. 2018 Jul 3;115(1):84-94. doi: 10.1016/j.bpj.2018.05.022. Biophys J. 2018. PMID: 29972814 Free PMC article.
-
Viral Membrane Fusion and the Transmembrane Domain.Viruses. 2020 Jun 27;12(7):693. doi: 10.3390/v12070693. Viruses. 2020. PMID: 32604992 Free PMC article. Review.
-
All-atom virus simulations.Curr Opin Virol. 2018 Aug;31:82-91. doi: 10.1016/j.coviro.2018.08.007. Epub 2018 Sep 1. Curr Opin Virol. 2018. PMID: 30181049 Free PMC article. Review.
-
Influence of interfacial tryptophan residues on an arginine-flanked transmembrane helix.Biochim Biophys Acta Biomembr. 2020 Feb 1;1862(2):183134. doi: 10.1016/j.bbamem.2019.183134. Epub 2019 Nov 16. Biochim Biophys Acta Biomembr. 2020. PMID: 31738898 Free PMC article.
-
Characterization of Lipid-Protein Interactions and Lipid-Mediated Modulation of Membrane Protein Function through Molecular Simulation.Chem Rev. 2019 May 8;119(9):6086-6161. doi: 10.1021/acs.chemrev.8b00608. Epub 2019 Apr 12. Chem Rev. 2019. PMID: 30978005 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

