Microscale thermophoresis provides insights into mechanism and thermodynamics of ribozyme catalysis
- PMID: 24448206
- PMCID: PMC3917984
- DOI: 10.4161/rna.27101
Microscale thermophoresis provides insights into mechanism and thermodynamics of ribozyme catalysis
Abstract
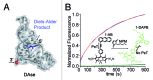
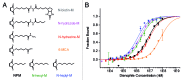
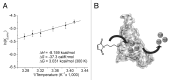
The analysis of binding interactions between small molecules and biopolymers is important for understanding biological processes. While fluorescence correlation spectroscopy (FCS) requires fluorescence labeling on the small molecule, which often interferes with binding, in microscale thermophoresis (MST) the label can be placed on the biopolymer. Ribozymes have not been analyzed by MST so far. The Diels-Alderase ribozyme (DAse) is a true catalyst, facilitating the Diels-Alder reaction between two free small substrates, anthracene dienes, and maleimide dienophiles. Despite high efforts, the determination of the dissociation constant (KD) of maleimide dienophiles to the DAse by FCS has been unsuccessful. Here, we determined the binding interactions of the DAse to its substrates and the Diels-Alder product using MST. The results supported a positive cooperativity for substrate binding to the DAse. By varying the temperature, we furthermore studied the thermodynamics of dienophile dissociation. The entropic contribution was found to be the energetic driving force for the binding of the dienophile to the DAse.
Keywords: Diels-Alder reaction; fluorescence correlation spectroscopy; microscale thermophoresis; ribozyme; thermodynamics.
Figures


Similar articles
-
Unravelling RNA-substrate interactions in a ribozyme-catalysed reaction using fluorescent turn-on probes.Chemistry. 2015 Apr 7;21(15):5864-71. doi: 10.1002/chem.201406512. Epub 2015 Mar 4. Chemistry. 2015. PMID: 25753253
-
Characterization of an RNA active site: interactions between a Diels-Alderase ribozyme and its substrates and products.J Am Chem Soc. 2002 Apr 3;124(13):3238-44. doi: 10.1021/ja0167405. J Am Chem Soc. 2002. PMID: 11916406
-
Stereoselective synthesis using immobilized Diels-Alderase ribozymes.Chembiochem. 2003 Oct 6;4(10):1089-92. doi: 10.1002/cbic.200300676. Chembiochem. 2003. PMID: 14523927
-
The structure and function of catalytic RNAs.Sci China C Life Sci. 2009 Mar;52(3):232-44. doi: 10.1007/s11427-009-0038-z. Epub 2009 Mar 18. Sci China C Life Sci. 2009. PMID: 19294348 Review.
-
Boron and Silicon-Substituted 1,3-Dienes and Dienophiles and Their Use in Diels-Alder Reactions.Molecules. 2020 Aug 16;25(16):3740. doi: 10.3390/molecules25163740. Molecules. 2020. PMID: 32824327 Free PMC article. Review.
Cited by
-
Target-Directed Approaches for Screening Small Molecules against RNA Targets.SLAS Discov. 2020 Sep;25(8):869-894. doi: 10.1177/2472555220922802. Epub 2020 May 18. SLAS Discov. 2020. PMID: 32419578 Free PMC article. Review.
-
Unveiling the druggable RNA targets and small molecule therapeutics.Bioorg Med Chem. 2019 May 15;27(10):2149-2165. doi: 10.1016/j.bmc.2019.03.057. Epub 2019 Mar 30. Bioorg Med Chem. 2019. PMID: 30981606 Free PMC article. Review.
-
Challenges with Simulating Modified RNA: Insights into Role and Reciprocity of Experimental and Computational Approaches.Genes (Basel). 2022 Mar 18;13(3):540. doi: 10.3390/genes13030540. Genes (Basel). 2022. PMID: 35328093 Free PMC article. Review.
-
Free Solution Assay Signal Modulation in Variable-Stem-Length Hairpin Aptamers.ACS Omega. 2020 May 14;5(20):11308-11313. doi: 10.1021/acsomega.9b04341. eCollection 2020 May 26. ACS Omega. 2020. PMID: 32478218 Free PMC article.
-
Measuring RNA-Ligand Interactions with Microscale Thermophoresis.Biochemistry. 2018 Aug 7;57(31):4638-4643. doi: 10.1021/acs.biochem.7b01141. Epub 2018 Jan 31. Biochemistry. 2018. PMID: 29327580 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
