Bronchial microbiome of severe COPD patients colonised by Pseudomonas aeruginosa
- PMID: 24449346
- PMCID: PMC4042013
- DOI: 10.1007/s10096-013-2044-0
Bronchial microbiome of severe COPD patients colonised by Pseudomonas aeruginosa
Abstract
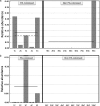
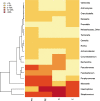
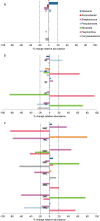
The bronchial microbiome in severe COPD during stability and exacerbation in patients chronically colonised by Pseudomonas aeruginosa (PA), has not been defined. Our objective was to determine the characteristics of the bronchial microbiome of severe COPD patients colonised and not colonised by P. aeruginosa and its changes during exacerbation. COPD patients with severe disease and frequent exacerbations were categorised according to chronic colonisation by P. aeruginosa. Sputum samples were obtained in stability and exacerbation, cultured, and analysed by 16S rRNA gene amplification and pyrosequencing. Sixteen patients were included, 5 of them showing chronic colonisation by P. aeruginosa. Pseudomonas genus had significantly higher relative abundance in stable colonised patients (p = 0.019), but no significant differences in biodiversity parameters were found between the two groups (Shannon, 3 (2-4) vs 3 (2-3), p = 0.699; Chao1, 124 (77-159) vs 140 (115-163), p = 0.364). In PA-colonised patients bronchial microbiome changed to a microbiome similar to non-PA-colonised patients during exacerbations. An increase in the relative abundance over 20 % during exacerbation was found for Streptococcus, Pseudomonas, Moraxella, Haemophilus, Neisseria, Achromobacter and Corynebacterium genera, which include recognised potentially pathogenic microorganisms, in 13 patients colonised and not colonised by P. aeruginosa with paired samples. These increases were not identified by culture in 5 out of 13 participants (38.5 %). Stable COPD patients with severe disease and PA-colonised showed a similar biodiversity to non-PA-colonised patients, with a higher relative abundance of Pseudomonas genus in bronchial secretions. Exacerbation in severe COPD patients showed the same microbial pattern, independently of previous colonisation by P. aeruginosa.
Figures

References
-
- Soler N, Torres A, Ewig S, Gonzalez J, Celis R, El-Ebiary M, Hernandez C, Rodriguez-Roisin R. Bronchial microbial patterns in severe exacerbations of chronic obstructive pulmonary disease (COPD) requiring mechanical ventilation. Am J Respir Crit Care Med. 1998;157(5 Pt 1):1498–1505. doi: 10.1164/ajrccm.157.5.9711044. - DOI - PubMed
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

