Direct cloning and refactoring of a silent lipopeptide biosynthetic gene cluster yields the antibiotic taromycin A
- PMID: 24449899
- PMCID: PMC3918841
- DOI: 10.1073/pnas.1319584111
Direct cloning and refactoring of a silent lipopeptide biosynthetic gene cluster yields the antibiotic taromycin A
Abstract
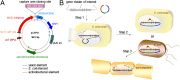
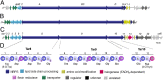
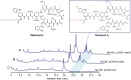
Recent developments in next-generation sequencing technologies have brought recognition of microbial genomes as a rich resource for novel natural product discovery. However, owing to the scarcity of efficient procedures to connect genes to molecules, only a small fraction of secondary metabolomes have been investigated to date. Transformation-associated recombination (TAR) cloning takes advantage of the natural in vivo homologous recombination of Saccharomyces cerevisiae to directly capture large genomic loci. Here we report a TAR-based genetic platform that allows us to directly clone, refactor, and heterologously express a silent biosynthetic pathway to yield a new antibiotic. With this method, which involves regulatory gene remodeling, we successfully expressed a 67-kb nonribosomal peptide synthetase biosynthetic gene cluster from the marine actinomycete Saccharomonospora sp. CNQ-490 and produced the dichlorinated lipopeptide antibiotic taromycin A in the model expression host Streptomyces coelicolor. The taromycin gene cluster (tar) is highly similar to the clinically approved antibiotic daptomycin from Streptomyces roseosporus, but has notable structural differences in three amino acid residues and the lipid side chain. With the activation of the tar gene cluster and production of taromycin A, this study highlights a unique "plug-and-play" approach to efficiently gaining access to orphan pathways that may open avenues for novel natural product discoveries and drug development.
Keywords: biosynthesis; drug discovery; genome mining.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Fleischmann RD, et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science. 1995;269(5223):496–512. - PubMed
-
- Bertelli C, Greub G. Rapid bacterial genome sequencing: Methods and applications in clinical microbiology. Clin Microbiol Infect. 2013;19(9):803–813. - PubMed
-
- Shendure J, Ji H. Next-generation DNA sequencing. Nat Biotechnol. 2008;26(10):1135–1145. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

