Zinc finger transcription factors displaced SREBP proteins as the major Sterol regulators during Saccharomycotina evolution
- PMID: 24453983
- PMCID: PMC3894159
- DOI: 10.1371/journal.pgen.1004076
Zinc finger transcription factors displaced SREBP proteins as the major Sterol regulators during Saccharomycotina evolution
Abstract
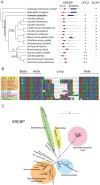
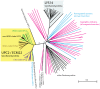
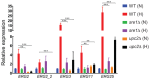
In most eukaryotes, including the majority of fungi, expression of sterol biosynthesis genes is regulated by Sterol-Regulatory Element Binding Proteins (SREBPs), which are basic helix-loop-helix transcription activators. However, in yeasts such as Saccharomyces cerevisiae and Candida albicans sterol synthesis is instead regulated by Upc2, an unrelated transcription factor with a Gal4-type zinc finger. The SREBPs in S. cerevisiae (Hms1) and C. albicans (Cph2) have lost a domain, are not major regulators of sterol synthesis, and instead regulate filamentous growth. We report here that rewiring of the sterol regulon, with Upc2 taking over from SREBP, likely occurred in the common ancestor of all Saccharomycotina. Yarrowia lipolytica, a deep-branching species, is the only genome known to contain intact and full-length orthologs of both SREBP (Sre1) and Upc2. Deleting YlUPC2, but not YlSRE1, confers susceptibility to azole drugs. Sterol levels are significantly reduced in the YlUPC2 deletion. RNA-seq analysis shows that hypoxic regulation of sterol synthesis genes in Y. lipolytica is predominantly mediated by Upc2. However, YlSre1 still retains a role in hypoxic regulation; growth of Y. lipolytica in hypoxic conditions is reduced in a Ylupc2 deletion and is abolished in a Ylsre1/Ylupc2 double deletion, and YlSre1 regulates sterol gene expression during hypoxia adaptation. We show that YlSRE1, and to a lesser extent YlUPC2, are required for switching from yeast to filamentous growth in hypoxia. Sre1 appears to have an ancestral role in the regulation of filamentation, which became decoupled from its role in sterol gene regulation by the arrival of Upc2 in the Saccharomycotina.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




References
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

