An efficient large-scale retroviral transduction method involving preloading the vector into a RetroNectin-coated bag with low-temperature shaking
- PMID: 24454964
- PMCID: PMC3893289
- DOI: 10.1371/journal.pone.0086275
An efficient large-scale retroviral transduction method involving preloading the vector into a RetroNectin-coated bag with low-temperature shaking
Abstract
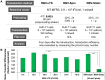
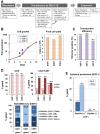
In retroviral vector-mediated gene transfer, transduction efficiency can be hampered by inhibitory molecules derived from the culture fluid of virus producer cell lines. To remove these inhibitory molecules to enable better gene transduction, we had previously developed a transduction method using a fibronectin fragment-coated vessel (i.e., the RetroNectin-bound virus transduction method). In the present study, we developed a method that combined RetroNectin-bound virus transduction with low-temperature shaking and applied this method in manufacturing autologous retroviral-engineered T cells for adoptive transfer gene therapy in a large-scale closed system. Retroviral vector was preloaded into a RetroNectin-coated bag and incubated at 4°C for 16 h on a reciprocating shaker at 50 rounds per minute. After the supernatant was removed, activated T cells were added to the bag. The bag transduction method has the advantage of increasing transduction efficiency, as simply flipping over the bag during gene transduction facilitates more efficient utilization of the retroviral vector adsorbed on the top and bottom surfaces of the bag. Finally, we performed validation runs of endoribonuclease MazF-modified CD4(+) T cell manufacturing for HIV-1 gene therapy and T cell receptor-modified T cell manufacturing for MAGE-A4 antigen-expressing cancer gene therapy and achieved over 200-fold (≥ 10(10)) and 100-fold (≥ 5 × 10(9)) expansion, respectively. In conclusion, we demonstrated that the large-scale closed transduction system is highly efficient for retroviral vector-based T cell manufacturing for adoptive transfer gene therapy, and this technology is expected to be amenable to automation and improve current clinical gene therapy protocols.
Conflict of interest statement
Figures



Similar articles
-
Influence of Retronectin-Mediated T-Cell Activation on Expansion and Phenotype of CD19-Specific Chimeric Antigen Receptor T Cells.Hum Gene Ther. 2018 Oct;29(10):1167-1182. doi: 10.1089/hum.2017.237. Epub 2018 Sep 5. Hum Gene Ther. 2018. PMID: 30024314
-
Retronectin-assisted retroviral transduction of primary human T lymphocytes under good manufacturing practice conditions: tissue culture bag critically determines cell yield.Cytotherapy. 2008;10(4):406-16. doi: 10.1080/14653240801982961. Cytotherapy. 2008. PMID: 18574773
-
Multifactorial optimization of gammaretroviral gene transfer into human T lymphocytes for clinical application.Hum Gene Ther. 2007 Dec;18(12):1253-60. doi: 10.1089/hum.2007.088. Hum Gene Ther. 2007. PMID: 18052719
-
Retroviral Vectors for Cancer Gene Therapy.Recent Results Cancer Res. 2016;209:17-35. doi: 10.1007/978-3-319-42934-2_2. Recent Results Cancer Res. 2016. PMID: 28101685 Review.
-
Protocol for making retroviral reprogramming factors.2012 Jun 10. In: StemBook [Internet]. Cambridge (MA): Harvard Stem Cell Institute; 2008–. 2012 Jun 10. In: StemBook [Internet]. Cambridge (MA): Harvard Stem Cell Institute; 2008–. PMID: 23658975 Free Books & Documents. Review.
Cited by
-
Autologous CD4 T Lymphocytes Modified with a Tat-Dependent, Virus-Specific Endoribonuclease Gene in HIV-Infected Individuals.Mol Ther. 2021 Feb 3;29(2):626-635. doi: 10.1016/j.ymthe.2020.11.007. Epub 2020 Nov 11. Mol Ther. 2021. PMID: 33186691 Free PMC article.
-
Microfluidic Transduction Harnesses Mass Transport Principles to Enhance Gene Transfer Efficiency.Mol Ther. 2017 Oct 4;25(10):2372-2382. doi: 10.1016/j.ymthe.2017.07.002. Epub 2017 Jul 8. Mol Ther. 2017. PMID: 28780274 Free PMC article.
-
A Rapid Cell Expansion Process for Production of Engineered Autologous CAR-T Cell Therapies.Hum Gene Ther Methods. 2016 Dec;27(6):209-218. doi: 10.1089/hgtb.2016.120. Hum Gene Ther Methods. 2016. PMID: 27897048 Free PMC article.
-
Artificial antigen-presenting cells: the booster for the obtaining of functional adoptive cells.Cell Mol Life Sci. 2024 Aug 31;81(1):378. doi: 10.1007/s00018-024-05412-y. Cell Mol Life Sci. 2024. PMID: 39215816 Free PMC article. Review.
-
miRNA-132-5p mediates a negative feedback regulation of IL-8 secretion through S100A8/A9 downregulation in neutrophil-like HL-60 cells.Front Immunol. 2024 Jan 16;14:1274378. doi: 10.3389/fimmu.2023.1274378. eCollection 2023. Front Immunol. 2024. PMID: 38292491 Free PMC article.
References
-
- Yoder MC, Williams DA (1995) Matrix molecule interactions with hematopoietic stem cells. Exp Hematol 23: 961–967. - PubMed
-
- Ruoslahti E (1988) Fibronectin and its receptors. Annu Rev Biochem 57: 375–413. - PubMed
-
- Hynes RO (1992) Integrins: versatility, modulation, and signaling in cell adhesion. Cell 69: 11–25. - PubMed
-
- Kimizuka F, Taguchi Y, Ohdate Y, Kawase Y, Shimojo T, et al. (1991) Production and characterization of functional domains of human fibronectin expressed in Escherichia coli . J Biochem 110: 284–291. - PubMed
-
- Hanenberg H, Xiao XL, Dilloo D, Hashino K, Kato I, et al. (1996) Colocalization of retrovirus and target cells on specific fibronectin fragments increases genetic transduction of mammalian cells. Nat Med 2: 876–882. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

