Sequential adaptive introgression of the mitochondrial genome in Drosophila yakuba and Drosophila santomea
- PMID: 24460929
- PMCID: PMC4260671
- DOI: 10.1111/mec.12678
Sequential adaptive introgression of the mitochondrial genome in Drosophila yakuba and Drosophila santomea
Abstract
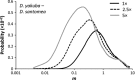
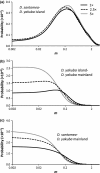
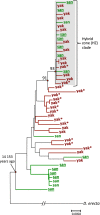
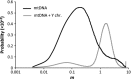
Interspecific hybridization provides the unique opportunity for species to tap into genetic variation present in a closely related species and potentially take advantage of beneficial alleles. It has become increasingly clear that when hybridization occurs, mitochondrial DNA (mtDNA) often crosses species boundaries, raising the possibility that it could serve as a recurrent target of natural selection and source of species' adaptations. Here we report the sequences of 46 complete mitochondrial genomes of Drosophila yakuba and Drosophila santomea, two sister species known to produce hybrids in nature (~3%). At least two independent events of mtDNA introgression are uncovered in this study, including an early invasion of the D. yakuba mitochondrial genome that fully replaced the D. santomea mtDNA native haplotypes and a more recent, ongoing event centred in the hybrid zone. Interestingly, this recent introgression event bears the signature of Darwinian natural selection, and the selective haplotype can be found at low frequency in Africa mainland populations of D. yakuba. We put forward the possibility that, because the effective population size of D. santomea is smaller than that of D. yakuba, the faster accumulation of mildly deleterious mutations associated with Muller's ratchet in the former species may have facilitated the replacement of the mutationally loaded mitochondrial genome of D. santomea by that of D. yakuba.
Keywords: gene flow; positive selection; recurrent adaptation; speciation.
© 2014 The Authors. Molecular Ecology Published by John Wiley & Sons Ltd.
Figures
Similar articles
-
Multilocus analysis of introgression between two sympatric sister species of Drosophila: Drosophila yakuba and D. santomea.Genetics. 2005 Sep;171(1):197-210. doi: 10.1534/genetics.104.033597. Epub 2005 Jun 18. Genetics. 2005. PMID: 15965264 Free PMC article.
-
The evolution of reproductive isolation in the Drosophila yakuba complex of species.J Evol Biol. 2015 Mar;28(3):557-75. doi: 10.1111/jeb.12588. J Evol Biol. 2015. PMID: 25611516
-
Gene flow between Drosophila yakuba and Drosophila santomea in subunit V of cytochrome c oxidase: A potential case of cytonuclear cointrogression.Evolution. 2015 Aug;69(8):1973-86. doi: 10.1111/evo.12718. Epub 2015 Aug 8. Evolution. 2015. PMID: 26155926 Free PMC article.
-
The Drosophila mitochondrial genome.Oxf Surv Eukaryot Genes. 1984;1:1-35. Oxf Surv Eukaryot Genes. 1984. PMID: 6400770 Review.
-
The on-again, off-again relationship between mitochondrial genomes and species boundaries.Mol Ecol. 2017 Apr;26(8):2212-2236. doi: 10.1111/mec.13959. Epub 2017 Jan 27. Mol Ecol. 2017. PMID: 27997046 Free PMC article. Review.
Cited by
-
Universal Mitochondrial Multi-Locus Sequence Analysis (mtMLSA) to Characterise Populations of Unanticipated Plant Pest Biosecurity Detections.Biology (Basel). 2022 Apr 24;11(5):654. doi: 10.3390/biology11050654. Biology (Basel). 2022. PMID: 35625382 Free PMC article.
-
Evolution of assortative mating following selective introgression of pigmentation genes between two Drosophila species.Ecol Evol. 2022 Apr 13;12(4):e8821. doi: 10.1002/ece3.8821. eCollection 2022 Apr. Ecol Evol. 2022. PMID: 35432924 Free PMC article.
-
Secondary reversion to sexual monomorphism associated with tissue-specific loss of doublesex expression.Evolution. 2022 Sep;76(9):2089-2104. doi: 10.1111/evo.14564. Epub 2022 Jul 23. Evolution. 2022. PMID: 35841603 Free PMC article.
-
The effects of allospecific mitochondrial genome on the fitness of northern redbelly dace (Chrosomus eos).Ecol Evol. 2018 Feb 19;8(6):3311-3321. doi: 10.1002/ece3.3922. eCollection 2018 Mar. Ecol Evol. 2018. PMID: 29607026 Free PMC article.
-
Wolbachia do not live by reproductive manipulation alone: infection polymorphism in Drosophila suzukii and D. subpulchrella.Mol Ecol. 2014 Oct;23(19):4871-85. doi: 10.1111/mec.12901. Epub 2014 Sep 18. Mol Ecol. 2014. PMID: 25156506 Free PMC article.
References
-
- Afonso JM, Pestano J, Hernandez M. Rapid isolation of mitochondrial DNA from Drosophila adults. Biochemical Genetics. 1988;26:381–386. - PubMed
-
- Anderson E. Introgressive Hybridization. New York: John Wiley & Sons; 1949.
-
- Arnold ML. Natural Hybridization and Evolution. Oxford: Oxford University Press; 1997.
-
- Arnold ML. Evolution through Genetic Exchange. Oxford: Oxford University Press; 2007.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

