Genomic analyses reveal broad impact of miR-137 on genes associated with malignant transformation and neuronal differentiation in glioblastoma cells
- PMID: 24465609
- PMCID: PMC3899048
- DOI: 10.1371/journal.pone.0085591
Genomic analyses reveal broad impact of miR-137 on genes associated with malignant transformation and neuronal differentiation in glioblastoma cells
Abstract
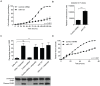
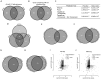
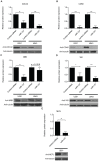
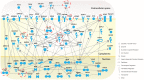
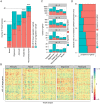
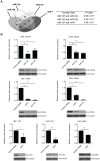
miR-137 plays critical roles in the nervous system and tumor development; an increase in its expression is required for neuronal differentiation while its reduction is implicated in gliomagenesis. To evaluate the potential of miR-137 in glioblastoma therapy, we conducted genome-wide target mapping in glioblastoma cells by measuring the level of association between PABP and mRNAs in cells transfected with miR-137 mimics vs. controls via RIPSeq. Impact on mRNA levels was also measured by RNASeq. By combining the results of both experimental approaches, 1468 genes were found to be negatively impacted by miR-137--among them, 595 (40%) contain miR-137 predicted sites. The most relevant targets include oncogenic proteins and key players in neurogenesis like c-KIT, YBX1, AKT2, CDC42, CDK6 and TGFβ2. Interestingly, we observed that several identified miR-137 targets are also predicted to be regulated by miR-124, miR-128 and miR-7, which are equally implicated in neuronal differentiation and gliomagenesis. We suggest that the concomitant increase of these four miRNAs in neuronal stem cells or their repression in tumor cells could produce a robust regulatory effect with major consequences to neuronal differentiation and tumorigenesis.
Conflict of interest statement
Figures
Similar articles
-
Involvement of miRNAs in the differentiation of human glioblastoma multiforme stem-like cells.PLoS One. 2013 Oct 14;8(10):e77098. doi: 10.1371/journal.pone.0077098. eCollection 2013. PLoS One. 2013. PMID: 24155920 Free PMC article.
-
Hypoxia-inducible miR-196a modulates glioblastoma cell proliferation and migration through complex regulation of NRAS.Cell Oncol (Dordr). 2021 Apr;44(2):433-451. doi: 10.1007/s13402-020-00580-y. Epub 2021 Jan 19. Cell Oncol (Dordr). 2021. PMID: 33469841
-
Antagonism between the RNA-binding protein Musashi1 and miR-137 and its potential impact on neurogenesis and glioblastoma development.RNA. 2019 Jul;25(7):768-782. doi: 10.1261/rna.069211.118. Epub 2019 Apr 19. RNA. 2019. PMID: 31004009 Free PMC article.
-
MiR-137: an important player in neural development and neoplastic transformation.Mol Psychiatry. 2017 Jan;22(1):44-55. doi: 10.1038/mp.2016.150. Epub 2016 Sep 13. Mol Psychiatry. 2017. PMID: 27620842 Free PMC article. Review.
-
Notch Signaling and MicroRNA: The Dynamic Duo Steering Between Neurogenesis and Glioblastomas.Cell Mol Biol (Noisy-le-grand). 2021 Aug 15;67(2):33-43. doi: 10.14715/cmb/2021.67.2.6. Cell Mol Biol (Noisy-le-grand). 2021. PMID: 34817376 Review.
Cited by
-
Role of microRNAs in glioblastoma.Oncotarget. 2021 Aug 17;12(17):1707-1723. doi: 10.18632/oncotarget.28039. eCollection 2021 Aug 17. Oncotarget. 2021. PMID: 34434499 Free PMC article.
-
MicroRNA Signature in Human Normal and Tumoral Neural Stem Cells.Int J Mol Sci. 2019 Aug 23;20(17):4123. doi: 10.3390/ijms20174123. Int J Mol Sci. 2019. PMID: 31450858 Free PMC article. Review.
-
MiR-137-3p Inhibits Colorectal Cancer Cell Migration by Regulating a KDM1A-Dependent Epithelial-Mesenchymal Transition.Dig Dis Sci. 2021 Jul;66(7):2272-2282. doi: 10.1007/s10620-020-06518-6. Epub 2020 Aug 4. Dig Dis Sci. 2021. PMID: 32749639
-
Synergistic effect of mesenchymal stem cell-derived extracellular vesicle and miR-137 alleviates autism-like behaviors by modulating the NF-κB pathway.J Transl Med. 2024 May 13;22(1):446. doi: 10.1186/s12967-024-05257-w. J Transl Med. 2024. PMID: 38741170 Free PMC article.
-
Identification of the Potential Prognostic Markers from the miRNA-lncRNA-mRNA Interactions for Metastatic Renal Cancer via Next-Generation Sequencing and Bioinformatics.Diagnostics (Basel). 2020 Apr 16;10(4):228. doi: 10.3390/diagnostics10040228. Diagnostics (Basel). 2020. PMID: 32316228 Free PMC article.
References
-
- Gonzalez-Gomez P, Sanchez P, Mira H (2011) MicroRNAs as regulators of neural stem cell-related pathways in glioblastoma multiforme. Mol Neurobiol 44: 235–249. - PubMed
-
- Perruisseau-Carrier C, Jurga M, Forraz N, McGuckin CP (2011) miRNAs stem cell reprogramming for neuronal induction and differentiation. Mol Neurobiol 43: 215–227. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

