Reconstruction of the carotenoid biosynthetic pathway of Cronobacter sakazakii BAA894 in Escherichia coli
- PMID: 24466219
- PMCID: PMC3900654
- DOI: 10.1371/journal.pone.0086739
Reconstruction of the carotenoid biosynthetic pathway of Cronobacter sakazakii BAA894 in Escherichia coli
Abstract
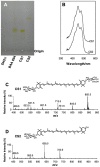
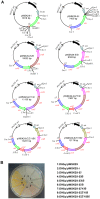
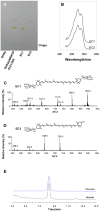
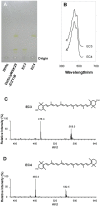
Cronobacter sakazakii could form yellow-pigmented colonies. However, the chemical structure and the biosynthetic pathway of the yellow pigments have not been identified. In this study, the yellow pigments of C. sakazakii BAA894 were purified and analyzed. The major components of the yellow pigments were confirmed as zeaxanthin-monoglycoside and zeaxanthin-diglycoside. A gene cluster containing seven genes responsible for the yellow pigmentation in C. sakazakii BAA894 was identified. The seven genes of C. sakazakii BAA894 or parts of them were reconstructed in a heterologous host Escherichia coli DH5α. The pigments formed in these E. coli strains were isolated and analyzed by thin layer chromatography, UV-visible spectroscopy, high performance liquid chromatography, and electron spray ionization-mass spectrometry. These redesigned E. coli strains could produce different carotenoids. E. coli strain expressing all the seven genes could produce zeaxanthin-monoglycoside and zeaxanthin-diglycoside; E. coli strains expressing parts of the seven genes could produce lycopene, β-carotene, cryptoxanthin or zeaxanthin. This study identified the gene cluster responsible for the yellow pigmentation in C. sakazakii BAA894.
Conflict of interest statement
Figures



Similar articles
-
Metabolic engineering of carotenoid accumulation in Escherichia coli by modulation of the isoprenoid precursor pool with expression of deoxyxylulose phosphate synthase.Appl Microbiol Biotechnol. 2000 Apr;53(4):396-400. doi: 10.1007/s002530051632. Appl Microbiol Biotechnol. 2000. PMID: 10803894
-
Genes involved in yellow pigmentation of Cronobacter sakazakii ES5 and influence of pigmentation on persistence and growth under environmental stress.Appl Environ Microbiol. 2010 Feb;76(4):1053-61. doi: 10.1128/AEM.01420-09. Epub 2009 Dec 28. Appl Environ Microbiol. 2010. PMID: 20038705 Free PMC article.
-
Cloning and characterization of Enterobacter sakazakii pigment genes and in situ spectroscopic analysis of the pigment.FEMS Microbiol Lett. 2006 Dec;265(2):244-8. doi: 10.1111/j.1574-6968.2006.00500.x. Epub 2006 Oct 27. FEMS Microbiol Lett. 2006. PMID: 17069626
-
Metabolic engineering for the production of carotenoids in non-carotenogenic bacteria and yeasts.J Biotechnol. 1997 Jan 3;59(3):169-81. doi: 10.1016/s0168-1656(97)00154-5. J Biotechnol. 1997. PMID: 9519479 Review.
-
Biological roles of fungal carotenoids.Curr Genet. 2015 Aug;61(3):309-24. doi: 10.1007/s00294-014-0454-x. Epub 2014 Oct 5. Curr Genet. 2015. PMID: 25284291 Review.
Cited by
-
A New Insight into the Bactericidal Mechanism of 405 nm Blue Light-Emitting-Diode against Dairy Sourced Cronobacter sakazakii.Foods. 2021 Aug 25;10(9):1996. doi: 10.3390/foods10091996. Foods. 2021. PMID: 34574108 Free PMC article.
-
The Global Response of Cronobacter sakazakii Cells to Amino Acid Deficiency.Front Microbiol. 2018 Aug 14;9:1875. doi: 10.3389/fmicb.2018.01875. eCollection 2018. Front Microbiol. 2018. PMID: 30154778 Free PMC article.
-
Synthetic biology and metabolic engineering for marine carotenoids: new opportunities and future prospects.Mar Drugs. 2014 Sep 17;12(9):4810-32. doi: 10.3390/md12094810. Mar Drugs. 2014. PMID: 25233369 Free PMC article. Review.
-
Complete Genome Sequence Analysis of Enterobacter sp. SA187, a Plant Multi-Stress Tolerance Promoting Endophytic Bacterium.Front Microbiol. 2017 Oct 20;8:2023. doi: 10.3389/fmicb.2017.02023. eCollection 2017. Front Microbiol. 2017. PMID: 29163376 Free PMC article.
-
Indigoidine biosynthesis triggered by the heavy metal-responsive transcription regulator: a visual whole-cell biosensor.Appl Microbiol Biotechnol. 2021 Aug;105(14-15):6087-6102. doi: 10.1007/s00253-021-11441-5. Epub 2021 Jul 22. Appl Microbiol Biotechnol. 2021. PMID: 34291315
References
-
- Cazzonelli CI, Pogson BJ (2010) Source to sink: regulation of carotenoid biosynthesis in plants. Trends Plant Sci 15: 266–274. - PubMed
-
- Higuera-Ciapara I, Felix-Valenzuela L, Goycoolea F (2006) Astaxanthin: a review of its chemistry and applications. Crit Rev Food Sci Nutr 46: 185–196. - PubMed
-
- Vachali P, Bhosale P, Bernstein PS (2012) Microbial carotenoids. Method Mol Biol 898: 41–59. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

