Critical roles for Rictor/Sin1 complexes in interferon-dependent gene transcription and generation of antiproliferative responses
- PMID: 24469448
- PMCID: PMC3945321
- DOI: 10.1074/jbc.M113.537852
Critical roles for Rictor/Sin1 complexes in interferon-dependent gene transcription and generation of antiproliferative responses
Abstract
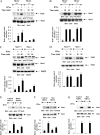
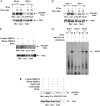
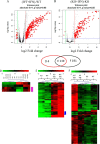
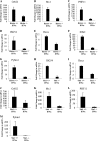
We provide evidence that type I IFN-induced STAT activation is diminished in cells with targeted disruption of the Rictor gene, whose protein product is a key element of mTOR complex 2. Our studies show that transient or stable knockdown of Rictor or Sin1 results in defects in activation of elements of the STAT pathway and reduced STAT-DNA binding complexes. This leads to decreased expression of several IFN-inducible genes that mediate important biological functions. Our studies also demonstrate that Rictor and Sin1 play essential roles in the generation of the suppressive effects of IFNα on malignant erythroid precursors from patients with myeloproliferative neoplasms. Altogether, these findings provide evidence for critical functions for Rictor/Sin1 complexes in type I IFN signaling and the generation of type I IFN antineoplastic responses.
Keywords: Akt PKB; Antiviral Agents; Cell Signaling; Cytokines/Interferon; Gene Regulation; Interferon; Signal Transduction.
Figures
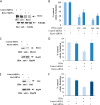
References
-
- Dunn G. P., Koebel C. M., Schreiber R. D. (2006) Interferons, immunity and cancer immunoediting. Nat. Rev. Immunol. 6, 836–848 - PubMed
-
- Hervas-Stubbs S., Perez-Gracia J. L., Rouzaut A., Sanmamed M. F., Le Bon A., Melero I. (2011) Direct effects of type I interferons on cells of the immune system. Clin. Cancer Res. 17, 2619–2627 - PubMed
-
- Platanias L. C. (2013) Interferons and their antitumor properties. J. Interferon Cytokine Res. 33, 143–144 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

