RNA design rules from a massive open laboratory
- PMID: 24469816
- PMCID: PMC3926058
- DOI: 10.1073/pnas.1313039111
RNA design rules from a massive open laboratory
Erratum in
-
Correction for Lee et al., RNA design rules from a massive open laboratory.Proc Natl Acad Sci U S A. 2015 Oct 27;112(43):E5900. doi: 10.1073/pnas.1518065112. Epub 2015 Oct 19. Proc Natl Acad Sci U S A. 2015. PMID: 26483501 Free PMC article. No abstract available.
-
Correction to Supporting Information for Lee et al., RNA design rules from a massive open laboratory.Proc Natl Acad Sci U S A. 2016 Jan 26;113(4):E490. doi: 10.1073/pnas.1525261113. Epub 2016 Jan 19. Proc Natl Acad Sci U S A. 2016. PMID: 26787869 Free PMC article. No abstract available.
Abstract
Self-assembling RNA molecules present compelling substrates for the rational interrogation and control of living systems. However, imperfect in silico models--even at the secondary structure level--hinder the design of new RNAs that function properly when synthesized. Here, we present a unique and potentially general approach to such empirical problems: the Massive Open Laboratory. The EteRNA project connects 37,000 enthusiasts to RNA design puzzles through an online interface. Uniquely, EteRNA participants not only manipulate simulated molecules but also control a remote experimental pipeline for high-throughput RNA synthesis and structure mapping. We show herein that the EteRNA community leveraged dozens of cycles of continuous wet laboratory feedback to learn strategies for solving in vitro RNA design problems on which automated methods fail. The top strategies--including several previously unrecognized negative design rules--were distilled by machine learning into an algorithm, EteRNABot. Over a rigorous 1-y testing phase, both the EteRNA community and EteRNABot significantly outperformed prior algorithms in a dozen RNA secondary structure design tests, including the creation of dendrimer-like structures and scaffolds for small molecule sensors. These results show that an online community can carry out large-scale experiments, hypothesis generation, and algorithm design to create practical advances in empirical science.
Keywords: RNA folding; citizen science; crowdsourcing; high-throughput experiments.
Figures
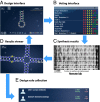



Similar articles
-
EternaBrain: Automated RNA design through move sets and strategies from an Internet-scale RNA videogame.PLoS Comput Biol. 2019 Jun 27;15(6):e1007059. doi: 10.1371/journal.pcbi.1007059. eCollection 2019 Jun. PLoS Comput Biol. 2019. PMID: 31247029 Free PMC article.
-
Principles for Predicting RNA Secondary Structure Design Difficulty.J Mol Biol. 2016 Feb 27;428(5 Pt A):748-757. doi: 10.1016/j.jmb.2015.11.013. Epub 2016 Feb 17. J Mol Biol. 2016. PMID: 26902426 Free PMC article.
-
Scientific rigor through videogames.Trends Biochem Sci. 2014 Nov;39(11):507-9. doi: 10.1016/j.tibs.2014.08.005. Epub 2014 Oct 6. Trends Biochem Sci. 2014. PMID: 25300714
-
Squaring theory with practice in RNA design.Curr Opin Struct Biol. 2012 Aug;22(4):457-66. doi: 10.1016/j.sbi.2012.06.003. Epub 2012 Jul 23. Curr Opin Struct Biol. 2012. PMID: 22832174 Review.
-
Recent advances in RNA folding.J Biotechnol. 2017 Nov 10;261:97-104. doi: 10.1016/j.jbiotec.2017.07.007. Epub 2017 Jul 8. J Biotechnol. 2017. PMID: 28690134 Review.
Cited by
-
Summary of the DREAM8 Parameter Estimation Challenge: Toward Parameter Identification for Whole-Cell Models.PLoS Comput Biol. 2015 May 28;11(5):e1004096. doi: 10.1371/journal.pcbi.1004096. eCollection 2015 May. PLoS Comput Biol. 2015. PMID: 26020786 Free PMC article.
-
Automated band annotation for RNA structure probing experiments with numerous capillary electrophoresis profiles.Bioinformatics. 2015 Sep 1;31(17):2808-15. doi: 10.1093/bioinformatics/btv282. Epub 2015 May 5. Bioinformatics. 2015. PMID: 25943472 Free PMC article.
-
Theoretical basis for stabilizing messenger RNA through secondary structure design.bioRxiv [Preprint]. 2021 Feb 19:2020.08.22.262931. doi: 10.1101/2020.08.22.262931. bioRxiv. 2021. Update in: Nucleic Acids Res. 2021 Oct 11;49(18):10604-10617. doi: 10.1093/nar/gkab764. PMID: 32869022 Free PMC article. Updated. Preprint.
-
A pipeline for computational design of novel RNA-like topologies.Nucleic Acids Res. 2018 Aug 21;46(14):7040-7051. doi: 10.1093/nar/gky524. Nucleic Acids Res. 2018. PMID: 30137633 Free PMC article.
-
RiboDraw: semiautomated two-dimensional drawing of RNA tertiary structure diagrams.NAR Genom Bioinform. 2021 Oct 14;3(4):lqab091. doi: 10.1093/nargab/lqab091. eCollection 2021 Dec. NAR Genom Bioinform. 2021. PMID: 34661102 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

