A novel computational biostatistics approach implies impaired dephosphorylation of growth factor receptors as associated with severity of autism
- PMID: 24473445
- PMCID: PMC3905234
- DOI: 10.1038/tp.2013.124
A novel computational biostatistics approach implies impaired dephosphorylation of growth factor receptors as associated with severity of autism
Abstract
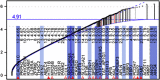
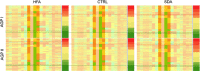
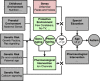
The prevalence of autism spectrum disorders (ASDs) has increased 20-fold over the past 50 years to >1% of US children. Although twin studies attest to a high degree of heritability, the genetic risk factors are still poorly understood. We analyzed data from two independent populations using u-statistics for genetically structured wide-locus data and added data from unrelated controls to explore epistasis. To account for systematic, but disease-unrelated differences in (non-randomized) genome-wide association studies (GWAS), a correlation between P-values and minor allele frequency with low granularity data and for conducting multiple tests in overlapping genetic regions, we present a novel study-specific criterion for 'genome-wide significance'. From recent results in a comorbid disease, childhood absence epilepsy, we had hypothesized that axonal guidance and calcium signaling are involved in autism as well. Enrichment of the results in both studies with related genes confirms this hypothesis. Additional ASD-specific variations identified in this study suggest protracted growth factor signaling as causing more severe forms of ASD. Another cluster of related genes suggests chloride and potassium ion channels as additional ASD-specific drug targets. The involvement of growth factors suggests the time of accelerated neuronal growth and pruning at 9-24 months of age as the period during which treatment with ion channel modulators would be most effective in preventing progression to more severe forms of autism. By extension, the same computational biostatistics approach could yield profound insights into the etiology of many common diseases from the genetic data collected over the last decade.
Figures



Comment in
-
Autism treatments proposed by clinical studies and human genetics are complementary.Transl Psychiatry. 2014 Jul 29;4(7):e415. doi: 10.1038/tp.2014.32. Transl Psychiatry. 2014. PMID: 25072320 Free PMC article. No abstract available.
Similar articles
-
Autism treatments proposed by clinical studies and human genetics are complementary.Transl Psychiatry. 2014 Jul 29;4(7):e415. doi: 10.1038/tp.2014.32. Transl Psychiatry. 2014. PMID: 25072320 Free PMC article. No abstract available.
-
Rare structural variation of synapse and neurotransmission genes in autism.Mol Psychiatry. 2012 Apr;17(4):402-11. doi: 10.1038/mp.2011.10. Epub 2011 Mar 1. Mol Psychiatry. 2012. PMID: 21358714 Free PMC article.
-
[Genetic analyses for identifying molecular mechanisms in autism spectrum disorders].Z Kinder Jugendpsychiatr Psychother. 2011 Mar;39(2):101-11. doi: 10.1024/1422-4917/a000096. Z Kinder Jugendpsychiatr Psychother. 2011. PMID: 21442598 Review. German.
-
Allowing for sex differences increases power in a GWAS of multiplex Autism families.Mol Psychiatry. 2012 Feb;17(2):215-22. doi: 10.1038/mp.2010.127. Epub 2010 Dec 14. Mol Psychiatry. 2012. PMID: 21151189
-
The genetic and neurobiologic compass points toward common signaling dysfunctions in autism spectrum disorders.J Clin Invest. 2009 Apr;119(4):747-54. doi: 10.1172/JCI37934. Epub 2009 Apr 1. J Clin Invest. 2009. PMID: 19339766 Free PMC article. Review.
Cited by
-
Expression profiling of disease progression in canine model of Duchenne muscular dystrophy.PLoS One. 2018 Mar 19;13(3):e0194485. doi: 10.1371/journal.pone.0194485. eCollection 2018. PLoS One. 2018. PMID: 29554127 Free PMC article.
-
Exploring the Contribution to ADHD of Genes Involved in Mendelian Disorders Presenting with Hyperactivity and/or Inattention.Genes (Basel). 2021 Dec 30;13(1):93. doi: 10.3390/genes13010093. Genes (Basel). 2021. PMID: 35052433 Free PMC article.
-
Mitochondrial Aspartate/Glutamate Carrier SLC25A12 and Autism Spectrum Disorder: a Meta-Analysis.Mol Neurobiol. 2016 Apr;53(3):1579-1588. doi: 10.1007/s12035-015-9116-3. Epub 2015 Feb 10. Mol Neurobiol. 2016. PMID: 25663199
-
The autism-associated MET receptor tyrosine kinase engages early neuronal growth mechanism and controls glutamatergic circuits development in the forebrain.Mol Psychiatry. 2016 Jul;21(7):925-35. doi: 10.1038/mp.2015.182. Epub 2016 Jan 5. Mol Psychiatry. 2016. PMID: 26728565 Free PMC article.
-
Pathway Network Analyses for Autism Reveal Multisystem Involvement, Major Overlaps with Other Diseases and Convergence upon MAPK and Calcium Signaling.PLoS One. 2016 Apr 7;11(4):e0153329. doi: 10.1371/journal.pone.0153329. eCollection 2016. PLoS One. 2016. PMID: 27055244 Free PMC article.
References
-
- Bailey A, Le Couteur A, Gottesman I, Bolton P, Simonoff E, Yuzda E, et al. Autism as a strongly genetic disorder: evidence from a British twin study. Psychol Med. 1995;25:63–77. - PubMed
-
- Devlin B, Scherer SW. Genetic architecture in autism spectrum disorder. Curr Opin Genet Dev. 2012;22:229–237. - PubMed
-
- Rutter M. Changing Concepts and Findings on Autism. J Autism Dev Disord. 2012;43:1749–1757. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- MH100182/MH/NIMH NIH HHS/United States
- R03 MH092618/MH/NIMH NIH HHS/United States
- MH086732/MH/NIMH NIH HHS/United States
- MH18268/MH/NIMH NIH HHS/United States
- P01 HD003008/HD/NICHD NIH HHS/United States
- UL1 RR024143/RR/NCRR NIH HHS/United States
- 8 UL1 TR000043/TR/NCATS NIH HHS/United States
- HD003008/HD/NICHD NIH HHS/United States
- R25DK092170-01A1/DK/NIDDK NIH HHS/United States
- UL1 TR000043/TR/NCATS NIH HHS/United States
- UL1 RR024139/RR/NCRR NIH HHS/United States
- T32 MH018268/MH/NIMH NIH HHS/United States
- R25 DK092170/DK/NIDDK NIH HHS/United States
- R01 MH100182/MH/NIMH NIH HHS/United States
- R03 MH086732/MH/NIMH NIH HHS/United States
- MH092618-01A1/MH/NIMH NIH HHS/United States
- 2 UL1 RR024143/RR/NCRR NIH HHS/United States
- UL1 TR000038/TR/NCATS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

