Enzyme recruitment and its role in metabolic expansion
- PMID: 24483367
- PMCID: PMC3985712
- DOI: 10.1021/bi401667f
Enzyme recruitment and its role in metabolic expansion
Abstract
Although more than 10(9) years have passed since the existence of the last universal common ancestor, proteins have yet to reach the limits of divergence. As a result, metabolic complexity is ever expanding. Identifying and understanding the mechanisms that drive and limit the divergence of protein sequence space impact not only evolutionary biologists investigating molecular evolution but also synthetic biologists seeking to design useful catalysts and engineer novel metabolic pathways. Investigations over the past 50 years indicate that the recruitment of enzymes for new functions is a key event in the acquisition of new metabolic capacity. In this review, we outline the genetic mechanisms that enable recruitment and summarize the present state of knowledge regarding the functional characteristics of extant catalysts that facilitate recruitment. We also highlight recent examples of enzyme recruitment, both from the historical record provided by phylogenetics and from enzyme evolution experiments. We conclude with a look to the future, which promises fruitful consequences from the convergence of molecular evolutionary theory, laboratory-directed evolution, and synthetic biology.
Figures
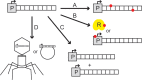

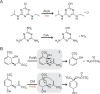
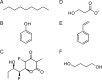
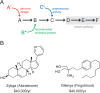
Similar articles
-
Directed evolution of enzymes and pathways for industrial biocatalysis.Curr Opin Biotechnol. 2002 Apr;13(2):104-10. doi: 10.1016/s0958-1669(02)00291-4. Curr Opin Biotechnol. 2002. PMID: 11950559 Review.
-
Dynamics and constraints of enzyme evolution.J Exp Zool B Mol Dev Evol. 2014 Nov;322(7):468-87. doi: 10.1002/jez.b.22562. Epub 2014 Feb 13. J Exp Zool B Mol Dev Evol. 2014. PMID: 24522979 Review.
-
Evolution of Enzyme Superfamilies: Comprehensive Exploration of Sequence-Function Relationships.Biochemistry. 2016 Nov 22;55(46):6375-6388. doi: 10.1021/acs.biochem.6b00723. Epub 2016 Nov 11. Biochemistry. 2016. PMID: 27802036 Review.
-
Directed Evolution of Protein Catalysts.Annu Rev Biochem. 2018 Jun 20;87:131-157. doi: 10.1146/annurev-biochem-062917-012034. Epub 2018 Mar 1. Annu Rev Biochem. 2018. PMID: 29494241 Review.
-
Metabolite-Enzyme Coevolution: From Single Enzymes to Metabolic Pathways and Networks.Annu Rev Biochem. 2018 Jun 20;87:187-216. doi: 10.1146/annurev-biochem-062917-012023. Annu Rev Biochem. 2018. PMID: 29925259 Review.
Cited by
-
Cell Survival Enabled by Leakage of a Labile Metabolic Intermediate.Mol Biol Evol. 2023 Mar 4;40(3):msad032. doi: 10.1093/molbev/msad032. Mol Biol Evol. 2023. PMID: 36788592 Free PMC article.
-
Emergence of terpene chemical communication in insects: Evolutionary recruitment of isoprenoid metabolism.Protein Sci. 2023 May;32(5):e4634. doi: 10.1002/pro.4634. Protein Sci. 2023. PMID: 36974623 Free PMC article.
-
Rapid bursts and slow declines: on the possible evolutionary trajectories of enzymes.J R Soc Interface. 2015 Jun 6;12(107):20150036. doi: 10.1098/rsif.2015.0036. J R Soc Interface. 2015. PMID: 25926697 Free PMC article. Review.
-
Molecular cloning, antiserum preparation and expression analysis during head regeneration of α-crystallin type heat shock protein in Hydra vulgaris.J Genet. 2018 Sep;97(4):911-924. J Genet. 2018. PMID: 30262703
-
Evolution of pollutant biodegradation.Appl Microbiol Biotechnol. 2025 Feb 4;109(1):36. doi: 10.1007/s00253-025-13418-0. Appl Microbiol Biotechnol. 2025. PMID: 39903283 Free PMC article. Review.
References
-
- Jensen R. A. (1976) Enzyme recruitment in evolution of new function. Annu. Rev. Microbiol. 30, 409–425. - PubMed
-
- Caetano-Anolles G.; Yafremava L. S.; Gee H.; Caetano-Anolles D.; Kim H. S.; Mittenthal J. E. (2009) The origin and evolution of modern metabolism. Int. J. Biochem. Cell Biol. 41, 285–297. - PubMed
-
- Gerlt J. A.; Babbitt P. C. (2001) Divergent evolution of enzymatic function: Mechanistically diverse superfamilies and functionally distinct suprafamilies. Annu. Rev. Biochem. 70, 209–246. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

