Characterization of mitochondrial haplogroups in a large population-based sample from the United States
- PMID: 24488180
- PMCID: PMC4113317
- DOI: 10.1007/s00439-014-1421-9
Characterization of mitochondrial haplogroups in a large population-based sample from the United States
Abstract
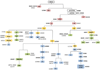
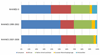
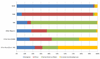
Mitochondrial DNA (mtDNA) haplogroups are valuable for investigations in forensic science, molecular anthropology, and human genetics. In this study, we developed a custom panel of 61 mtDNA markers for high-throughput classification of European, African, and Native American/Asian mitochondrial haplogroup lineages. Using these mtDNA markers, we constructed a mitochondrial haplogroup classification tree and classified 18,832 participants from the National Health and Nutrition Examination Surveys (NHANES). To our knowledge, this is the largest study to date characterizing mitochondrial haplogroups in a population-based sample from the United States, and the first study characterizing mitochondrial haplogroup distributions in self-identified Mexican Americans separately from Hispanic Americans of other descent. We observed clear differences in the distribution of maternal genetic ancestry consistent with proposed admixture models for these subpopulations, underscoring the genetic heterogeneity of the United States Hispanic population. The mitochondrial haplogroup distributions in the other self-identified racial/ethnic groups within NHANES were largely comparable to previous studies. Mitochondrial haplogroup classification was highly concordant with self-identified race/ethnicity (SIRE) in non-Hispanic whites (94.8 %), but was considerably lower in admixed populations including non-Hispanic blacks (88.3 %), Mexican Americans (81.8 %), and other Hispanics (61.6 %), suggesting SIRE does not accurately reflect maternal genetic ancestry, particularly in populations with greater proportions of admixture. Thus, it is important to consider inconsistencies between SIRE and genetic ancestry when performing genetic association studies. The mitochondrial haplogroup data that we have generated, coupled with the epidemiologic variables in NHANES, is a valuable resource for future studies investigating the contribution of mtDNA variation to human health and disease.
Figures
Similar articles
-
Estimates of continental ancestry vary widely among individuals with the same mtDNA haplogroup.Am J Hum Genet. 2015 Feb 5;96(2):183-93. doi: 10.1016/j.ajhg.2014.12.015. Epub 2015 Jan 22. Am J Hum Genet. 2015. PMID: 25620206 Free PMC article.
-
Mitochondrial variation and the risk of age-related macular degeneration across diverse populations.Pac Symp Biocomput. 2015:243-54. Pac Symp Biocomput. 2015. PMID: 25592585 Free PMC article.
-
Evaluation of variation in control region sequences for Hispanic individuals in the SWGDAM mtDNA data set.J Forensic Sci. 2006 May;51(3):566-73. doi: 10.1111/j.1556-4029.2006.00136.x. J Forensic Sci. 2006. PMID: 16696703
-
Patterns of mtDNA diversity in northwestern North America.Hum Biol. 2004 Feb;76(1):33-54. doi: 10.1353/hub.2004.0023. Hum Biol. 2004. PMID: 15222679 Review.
-
Recent progress in mitochondrial DNA analysis.Leg Med (Tokyo). 2005 Jul;7(4):259-62. doi: 10.1016/j.legalmed.2005.01.005. Leg Med (Tokyo). 2005. PMID: 15939655 Review.
Cited by
-
Mitochondrial Function in Health and Disease: Responses to Helicobacter pylori Metabolism and Impact in Gastric Cancer Development.Curr Top Microbiol Immunol. 2023;444:53-81. doi: 10.1007/978-3-031-47331-9_3. Curr Top Microbiol Immunol. 2023. PMID: 38231215
-
Mitochondria and Alzheimer's Disease: the Role of Mitochondrial Genetic Variation.Curr Genet Med Rep. 2018;6(1):1-10. doi: 10.1007/s40142-018-0132-2. Epub 2018 Mar 1. Curr Genet Med Rep. 2018. PMID: 29564191 Free PMC article. Review.
-
Haplotype diversity in mitochondrial DNA hypervariable region in a population of southeastern Brazil.Int J Legal Med. 2014 Jul;128(4):589-93. doi: 10.1007/s00414-014-1023-z. Epub 2014 May 21. Int J Legal Med. 2014. PMID: 24846100
-
Altered Retrograde Signaling Patterns in Breast Cancer Cells Cybrids with H and J Mitochondrial DNA Haplogroups.Int J Mol Sci. 2022 Jun 15;23(12):6687. doi: 10.3390/ijms23126687. Int J Mol Sci. 2022. PMID: 35743133 Free PMC article.
-
Natural Barcodes for Longitudinal Single Cell Tracking of Leukemic and Immune Cell Dynamics.Front Immunol. 2022 Jan 3;12:788891. doi: 10.3389/fimmu.2021.788891. eCollection 2021. Front Immunol. 2022. PMID: 35046946 Free PMC article. Review.
References
-
- Allard MW, Miller K, Wilson M, Monson K, Budowle B. Characterization of the Caucasian haplogroups present in the SWGDAM forensic mtDNA dataset for 1771 human control region sequences. Scientific Working Group on DNA Analysis Methods. J Forensic Sci. 2002;47:1215–1223. - PubMed
-
- Allard MW, Polanskey D, Miller K, Wilson MR, Monson KL, Budowle B. Characterization of human control region sequences of the African American SWGDAM forensic mtDNA data set. Forensic Sci Int. 2005;148:169–179. - PubMed
-
- Allard MW, Polanskey D, Wilson MR, Monson KL, Budowle B. Evaluation of Variation in Control Region Sequences for Hispanic Individuals in the SWGDAM mtDNA Data Set. J Forensic Sci. 2006;51:566–573. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

