Structure of the chicken CD3εδ/γ heterodimer and its assembly with the αβT cell receptor
- PMID: 24488493
- PMCID: PMC3961652
- DOI: 10.1074/jbc.M113.544965
Structure of the chicken CD3εδ/γ heterodimer and its assembly with the αβT cell receptor
Abstract
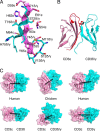
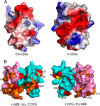
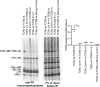
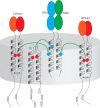
In mammals, the αβT cell receptor (TCR) signaling complex is composed of a TCRαβ heterodimer that is noncovalently coupled to three dimeric signaling molecules, CD3εδ, CD3εγ, and CD3ζζ. The nature of the TCR signaling complex and subunit arrangement in different species remains unclear however. Here we present a structural and biochemical analysis of the more primitive ancestral form of the TCR signaling complex found in chickens. In contrast to mammals, chickens do not express separate CD3δ and CD3γ chains but instead encode a single hybrid chain, termed CD3δ/γ, that is capable of pairing with CD3ε. The NMR structure of the chicken CD3εδ/γ heterodimer revealed a unique dimer interface that results in a heterodimer with considerable deviation from the distinct side-by-side architecture found in human and murine CD3εδ and CD3εγ. The chicken CD3εδ/γ heterodimer also contains a unique molecular surface, with the vast majority of surface-exposed, nonconserved residues being clustered to a single face of the heterodimer. Using an in vitro biochemical assay, we demonstrate that CD3εδ/γ can assemble with both chicken TCRα and TCRβ via conserved polar transmembrane sites. Moreover, analogous to the human TCR signaling complex, the presence of two copies of CD3εδ/γ is required for ζζ assembly. These data provide insight into the evolution of this critical receptor signaling apparatus.
Keywords: Cell Signaling; Immunology; Protein Assembly; Protein Evolution; Protein Structure.
Figures




Similar articles
-
Solution structure of the CD3epsilondelta ectodomain and comparison with CD3epsilongamma as a basis for modeling T cell receptor topology and signaling.Proc Natl Acad Sci U S A. 2004 Nov 30;101(48):16867-72. doi: 10.1073/pnas.0407576101. Epub 2004 Nov 19. Proc Natl Acad Sci U S A. 2004. PMID: 15557001 Free PMC article.
-
Expression and characterization of recombinant soluble human CD3 molecules: presentation of antigenic epitopes defined on the native TCR-CD3 complex.Int Immunol. 2002 Apr;14(4):389-400. doi: 10.1093/intimm/14.4.389. Int Immunol. 2002. PMID: 11934875
-
Evolution of T cell receptor (TCR) alpha beta heterodimer assembly with the CD3 complex.Eur J Immunol. 2001 Dec;31(12):3798-805. doi: 10.1002/1521-4141(200112)31:12<3798::aid-immu3798>3.0.co;2-z. Eur J Immunol. 2001. PMID: 11745401
-
The structural basis of αβ T-lineage immune recognition: TCR docking topologies, mechanotransduction, and co-receptor function.Immunol Rev. 2012 Nov;250(1):102-19. doi: 10.1111/j.1600-065X.2012.01161.x. Immunol Rev. 2012. PMID: 23046125 Free PMC article. Review.
-
Some hints concerning the shape of T-cell receptor structures.Scand J Immunol. 2002 Feb;55(2):111-8. doi: 10.1046/j.1365-3083.2002.01044.x. Scand J Immunol. 2002. PMID: 11896927 Review.
Cited by
-
Contribution of the TCRβ Repertoire to Marek's Disease Genetic Resistance in the Chicken.Viruses. 2023 Feb 22;15(3):607. doi: 10.3390/v15030607. Viruses. 2023. PMID: 36992316 Free PMC article.
-
Human γδ T cells augment antigen presentation in Listeria Monocytogenes infection.Mol Med. 2016 Dec;22:737-746. doi: 10.2119/molmed.2015.00214. Epub 2016 Sep 19. Mol Med. 2016. PMID: 27652377 Free PMC article.
-
Molecular architecture of the αβ T cell receptor-CD3 complex.Proc Natl Acad Sci U S A. 2014 Dec 9;111(49):17576-81. doi: 10.1073/pnas.1420936111. Epub 2014 Nov 24. Proc Natl Acad Sci U S A. 2014. PMID: 25422432 Free PMC article.
-
Structural and Biophysical Insights into the TCRαβ Complex in Chickens.iScience. 2020 Nov 20;23(12):101828. doi: 10.1016/j.isci.2020.101828. eCollection 2020 Dec 18. iScience. 2020. PMID: 33305184 Free PMC article.
-
The Chicken Embryo Model: A Novel and Relevant Model for Immune-Based Studies.Front Immunol. 2021 Nov 19;12:791081. doi: 10.3389/fimmu.2021.791081. eCollection 2021. Front Immunol. 2021. PMID: 34868080 Free PMC article. Review.
References
-
- Schamel W. W., Alarcón B. (2013) Organization of the resting TCR in nanoscale oligomers. Immunol. Rev. 251, 13–20 - PubMed
-
- Birkinshaw R. W., Kjer-Nielsen L., Eckle S. B. G., McCluskey J., Rossjohn J. (2014) MAITs, MR1 and vitamin B metabolites. Curr. Opin. Immunol. 26, 7–13 - PubMed
-
- Eckle S. B., Turner S. J., Rossjohn J., McCluskey J. (2013) Predisposed αβT cell antigen receptor recognition of MHC and MHC-I like molecules? Curr. Opin. Immunol. 25, 653–659 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

