SNP discovery in the transcriptome of white Pacific shrimp Litopenaeus vannamei by next generation sequencing
- PMID: 24498047
- PMCID: PMC3907553
- DOI: 10.1371/journal.pone.0087218
SNP discovery in the transcriptome of white Pacific shrimp Litopenaeus vannamei by next generation sequencing
Abstract
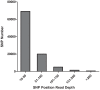
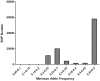
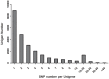
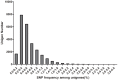
The application of next generation sequencing technology has greatly facilitated high throughput single nucleotide polymorphism (SNP) discovery and genotyping in genetic research. In the present study, SNPs were discovered based on two transcriptomes of Litopenaeus vannamei (L. vannamei) generated from Illumina sequencing platform HiSeq 2000. One transcriptome of L. vannamei was obtained through sequencing on the RNA from larvae at mysis stage and its reference sequence was de novo assembled. The data from another transcriptome were downloaded from NCBI and the reads of the two transcriptomes were mapped separately to the assembled reference by BWA. SNP calling was performed using SAMtools. A total of 58,717 and 36,277 SNPs with high quality were predicted from the two transcriptomes, respectively. SNP calling was also performed using the reads of two transcriptomes together, and a total of 96,040 SNPs with high quality were predicted. Among these 96,040 SNPs, 5,242 and 29,129 were predicted as non-synonymous and synonymous SNPs respectively. Characterization analysis of the predicted SNPs in L. vannamei showed that the estimated SNP frequency was 0.21% (one SNP per 476 bp) and the estimated ratio for transition to transversion was 2.0. Fifty SNPs were randomly selected for validation by Sanger sequencing after PCR amplification and 76% of SNPs were confirmed, which indicated that the SNPs predicted in this study were reliable. These SNPs will be very useful for genetic study in L. vannamei, especially for the high density linkage map construction and genome-wide association studies.
Conflict of interest statement
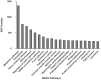
Figures


Similar articles
-
Analysis of Litopenaeus vannamei transcriptome using the next-generation DNA sequencing technique.PLoS One. 2012;7(10):e47442. doi: 10.1371/journal.pone.0047442. Epub 2012 Oct 11. PLoS One. 2012. PMID: 23071809 Free PMC article.
-
Identification of SNP markers associated with tolerance to ammonia toxicity by selective genotyping from de novo assembled transcriptome in Litopenaeus vannamei.Fish Shellfish Immunol. 2018 Feb;73:158-166. doi: 10.1016/j.fsi.2017.12.005. Epub 2017 Dec 5. Fish Shellfish Immunol. 2018. PMID: 29208499
-
Genome survey and high-density genetic map construction provide genomic and genetic resources for the Pacific White Shrimp Litopenaeus vannamei.Sci Rep. 2015 Oct 27;5:15612. doi: 10.1038/srep15612. Sci Rep. 2015. PMID: 26503227 Free PMC article.
-
Sex-Specific Transcriptome Sequencing of Zoea I Larvae and Identification of Sex-Linked Genes Using Bulked Segregant Analysis in Pacific White Shrimp Litopenaeus vannamei.Mar Biotechnol (NY). 2020 Jun;22(3):423-432. doi: 10.1007/s10126-020-09962-7. Epub 2020 Apr 12. Mar Biotechnol (NY). 2020. PMID: 32281012
-
Genome-wide SNP discovery from transcriptome of four common carp strains.PLoS One. 2012;7(10):e48140. doi: 10.1371/journal.pone.0048140. Epub 2012 Oct 26. PLoS One. 2012. PMID: 23110192 Free PMC article.
Cited by
-
Genomic signatures of artificial selection in fecundity of Pacific white shrimp, Penaeus vannamei.Front Genet. 2022 Aug 29;13:929889. doi: 10.3389/fgene.2022.929889. eCollection 2022. Front Genet. 2022. PMID: 36105098 Free PMC article.
-
Comparative Analysis of Root Transcriptome Reveals Candidate Genes and Expression Divergence of Homoeologous Genes in Response to Water Stress in Wheat.Plants (Basel). 2020 May 7;9(5):596. doi: 10.3390/plants9050596. Plants (Basel). 2020. PMID: 32392904 Free PMC article.
-
Litopenaeus vannamei Transcriptome Profile of Populations Evaluated for Growth Performance and Exposed to White Spot Syndrome Virus (WSSV).Front Genet. 2018 Apr 10;9:120. doi: 10.3389/fgene.2018.00120. eCollection 2018. Front Genet. 2018. PMID: 29692800 Free PMC article. No abstract available.
-
Immunomics of the koala (Phascolarctos cinereus).Immunogenetics. 2015 Jun;67(5-6):305-21. doi: 10.1007/s00251-015-0833-6. Epub 2015 Mar 13. Immunogenetics. 2015. PMID: 25761531
-
Identification of SNPs potentially related to immune responses and growth performance in Litopenaeus vannamei by RNA-seq analyses.PeerJ. 2018 Jul 4;6:e5154. doi: 10.7717/peerj.5154. eCollection 2018. PeerJ. 2018. PMID: 30013834 Free PMC article.
References
-
- Zhang L, Yang C, Zhang Y, Li L, Zhang X, et al. (2006) A genetic linkage map of Pacific white shrimp (Litopenaeus vannamei): sex-linked microsatellite markers and high recombination rates. Genetica 131: 37–49. - PubMed
-
- Alcivar-Warren A, Meehan-Meola D, Park SW, Xu Z, Delaney M, et al. (2007) ShrimpMap: A low-density, microsatellite-based linkage map of the pacific whiteleg shrimp, Litopenaeus vannamei: Identification of sex-linked markers in linkage group 4. Jounal of Shellfish Research 26: 1259–1277.
-
- Du ZQ, Ciobanu DC, Onteru SK, Gorbach D, Mileham AJ, et al. (2010) A gene-based SNP linkage map for pacific white shrimp, Litopenaeus vannamei . Animal Genetics 41: 286–294. - PubMed
-
- Perez F, Erazo C, Zhinaula M, Volckaert F, Calderon J (2004) A sex-specific linkage map of the white shrimp Penaeus (Litopenaeus) vannamei based on AFLP markers. Aquaculture 242: 105–118.
-
- Glenn KL, Grapes L, Suwanasopee T, Harris DL, Li Y, et al. (2005) SNP analysis of AMY2 and CTSL genes in Litopenaeus vannamei and Penaeus monodon shrimp. Animal Genetics 36: 235–236. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

