Shared gene structures and clusters of mutually exclusive spliced exons within the metazoan muscle myosin heavy chain genes
- PMID: 24498429
- PMCID: PMC3912159
- DOI: 10.1371/journal.pone.0088111
Shared gene structures and clusters of mutually exclusive spliced exons within the metazoan muscle myosin heavy chain genes
Abstract
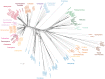
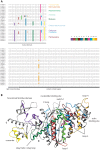
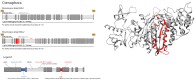
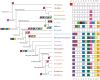
Multicellular animals possess two to three different types of muscle tissues. Striated muscles have considerable ultrastructural similarity and contain a core set of proteins including the muscle myosin heavy chain (Mhc) protein. The ATPase activity of this myosin motor protein largely dictates muscle performance at the molecular level. Two different solutions to adjusting myosin properties to different muscle subtypes have been identified so far: Vertebrates and nematodes contain many independent differentially expressed Mhc genes while arthropods have single Mhc genes with clusters of mutually exclusive spliced exons (MXEs). The availability of hundreds of metazoan genomes now allowed us to study whether the ancient bilateria already contained MXEs, how MXE complexity subsequently evolved, and whether additional scenarios to control contractile properties in different muscles could be proposed, By reconstructing the Mhc genes from 116 metazoans we showed that all intron positions within the motor domain coding regions are conserved in all bilateria analysed. The last common ancestor of the bilateria already contained a cluster of MXEs coding for part of the loop-2 actin-binding sequence. Subsequently the protostomes and later the arthropods gained many further clusters while MXEs got completely lost independently in several branches (vertebrates and nematodes) and species (for example the annelid Helobdella robusta and the salmon louse Lepeophtheirus salmonis). Several bilateria have been found to encode multiple Mhc genes that might all or in part contain clusters of MXEs. Notable examples are a cluster of six tandemly arrayed Mhc genes, of which two contain MXEs, in the owl limpet Lottia gigantea and four Mhc genes with three encoding MXEs in the predatory mite Metaseiulus occidentalis. Our analysis showed that similar solutions to provide different myosin isoforms (multiple genes or clusters of MXEs or both) have independently been developed several times within bilaterian evolution.
Conflict of interest statement
Figures




Similar articles
-
Comparative genomic analysis of the arthropod muscle myosin heavy chain genes allows ancestral gene reconstruction and reveals a new type of 'partially' processed pseudogene.BMC Mol Biol. 2008 Feb 6;9:21. doi: 10.1186/1471-2199-9-21. BMC Mol Biol. 2008. PMID: 18254963 Free PMC article.
-
Expansion of the mutually exclusive spliced exome in Drosophila.Nat Commun. 2013;4:2460. doi: 10.1038/ncomms3460. Nat Commun. 2013. PMID: 24025855
-
Functional domains of the Drosophila melanogaster muscle myosin heavy-chain gene are encoded by alternatively spliced exons.Mol Cell Biol. 1989 Jul;9(7):2957-74. doi: 10.1128/mcb.9.7.2957-2974.1989. Mol Cell Biol. 1989. PMID: 2506434 Free PMC article.
-
Alternative splicing of mutually exclusive exons--a review.Biosystems. 2013 Oct;114(1):31-8. doi: 10.1016/j.biosystems.2013.07.003. Epub 2013 Jul 11. Biosystems. 2013. PMID: 23850531 Review.
-
Postembryonic expression of the myosin heavy chain genes in the limb, tail, and heart muscles of metamorphosing amphibian tadpoles.Microsc Res Tech. 2000 Sep 15;50(6):458-72. doi: 10.1002/1097-0029(20000915)50:6<458::AID-JEMT4>3.0.CO;2-V. Microsc Res Tech. 2000. PMID: 10998636 Review.
Cited by
-
Myosin II sequences for Lethocerus indicus.J Muscle Res Cell Motil. 2017 Apr;38(2):193-200. doi: 10.1007/s10974-017-9476-6. Epub 2017 Jul 13. J Muscle Res Cell Motil. 2017. PMID: 28707142 Free PMC article.
-
Genome Sequencing of the Phytoseiid Predatory Mite Metaseiulus occidentalis Reveals Completely Atomized Hox Genes and Superdynamic Intron Evolution.Genome Biol Evol. 2016 Jun 27;8(6):1762-75. doi: 10.1093/gbe/evw048. Genome Biol Evol. 2016. PMID: 26951779 Free PMC article.
-
Genome-Wide Identification and Expression Profiles of Myosin Genes in the Pacific White Shrimp, Litopenaeus vannamei.Front Physiol. 2019 May 21;10:610. doi: 10.3389/fphys.2019.00610. eCollection 2019. Front Physiol. 2019. PMID: 31178751 Free PMC article.
-
Myosin repertoire expansion coincides with eukaryotic diversification in the Mesoproterozoic era.BMC Evol Biol. 2017 Sep 4;17(1):211. doi: 10.1186/s12862-017-1056-2. BMC Evol Biol. 2017. PMID: 28870165 Free PMC article.
-
Spaced words and kmacs: fast alignment-free sequence comparison based on inexact word matches.Nucleic Acids Res. 2014 Jul;42(Web Server issue):W7-11. doi: 10.1093/nar/gku398. Epub 2014 May 14. Nucleic Acids Res. 2014. PMID: 24829447 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

