Relative roles of GM1 ganglioside, N-acylneuraminic acids, and α2β1 integrin in mediating rotavirus infection
- PMID: 24501414
- PMCID: PMC3993774
- DOI: 10.1128/JVI.03431-13
Relative roles of GM1 ganglioside, N-acylneuraminic acids, and α2β1 integrin in mediating rotavirus infection
Abstract
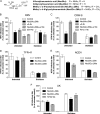
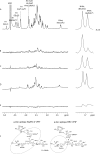
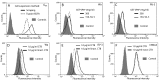
N-acetyl- and N-glycolylneuraminic acids (Sia) and α2β1 integrin are frequently used by rotaviruses as cellular receptors through recognition by virion spike protein VP4. The VP4 subunit VP8*, derived from Wa rotavirus, binds the internal N-acetylneuraminic acid on ganglioside GM1. Wa infection is increased by enhanced internal Sia access following terminal Sia removal from main glycan chains with sialidase. The GM1 ligand cholera toxin B (CTB) reduces Wa infectivity. Here, we found sialidase treatment increased cellular GM1 availability and the infectivity of several other human (including RV-3) and animal rotaviruses, typically rendering them susceptible to methyl α-d-N-acetylneuraminide treatment, but did not alter α2β1 usage. CTB reduced the infectivity of these viruses. Aceramido-GM1 inhibited Wa and RV-3 infectivity in untreated and sialidase-treated cells, and GM1 supplementation increased their infectivity, demonstrating the importance of GM1 for infection. Wa recognition of α2β1 and internal Sia were at least partially independent. Rotavirus usage of GM1 was mapped to VP4 using virus reassortants, and RV-3 VP8* bound aceramido-GM1 by saturation transfer difference nuclear magnetic resonance (STD NMR). Most rotaviruses recognizing terminal Sia did not use GM1, including RRV. RRV VP8* interacted minimally with aceramido-GM1 by STD NMR. Unusually, TFR-41 rotavirus infectivity depended upon terminal Sia and GM1. Competition of CTB, Sia, and/or aceramido-GM1 with cell binding by VP8* from representative rotaviruses showed that rotavirus Sia and GM1 preferences resulted from VP8*-cell binding. Our major finding is that infection by human rotaviruses of commonly occurring VP4 serotypes involves VP8* binding to cell surface GM1 glycan, typically including the internal N-acetylneuraminic acid.
Importance: Rotaviruses, the major cause of severe infantile gastroenteritis, recognize cell surface receptors through virus spike protein VP4. Several animal rotaviruses are known to bind sialic acids at the termini of main carbohydrate chains. Conversely, only a single human rotavirus is known to bind sialic acid. Interestingly, VP4 of this rotavirus bound to sialic acid that forms a branch on the main carbohydrate chain of the GM1 ganglioside. Here, we use several techniques to demonstrate that other human rotaviruses exhibit similar GM1 usage properties. Furthermore, binding by VP4 to cell surface GM1, involving branched sialic acid recognition, is shown to facilitate infection. In contrast, most animal rotaviruses that bind terminal sialic acids did not utilize GM1 for VP4 cell binding or infection. These studies support a significant role for GM1 in mediating host cell invasion by human rotaviruses.
Figures



Similar articles
-
Rotavirus spike protein VP5* binds alpha2beta1 integrin on the cell surface and competes with virus for cell binding and infectivity.J Gen Virol. 2006 May;87(Pt 5):1275-1283. doi: 10.1099/vir.0.81580-0. J Gen Virol. 2006. PMID: 16603530
-
Structural basis of rotavirus strain preference toward N-acetyl- or N-glycolylneuraminic acid-containing receptors.J Virol. 2012 Dec;86(24):13456-66. doi: 10.1128/JVI.06975-11. Epub 2012 Oct 3. J Virol. 2012. PMID: 23035213 Free PMC article.
-
Cell attachment protein VP8* of a human rotavirus specifically interacts with A-type histo-blood group antigen.Nature. 2012 Apr 15;485(7397):256-9. doi: 10.1038/nature10996. Nature. 2012. PMID: 22504179 Free PMC article.
-
Molecular biology of rotavirus cell entry.Arch Med Res. 2002 Jul-Aug;33(4):356-61. doi: 10.1016/s0188-4409(02)00374-0. Arch Med Res. 2002. PMID: 12234525 Review.
-
Carbohydrate recognition by rotaviruses.J Struct Funct Genomics. 2014 Sep;15(3):101-6. doi: 10.1007/s10969-013-9167-5. Epub 2013 Nov 19. J Struct Funct Genomics. 2014. PMID: 24248423 Review.
Cited by
-
Glycosphingolipid-Protein Interaction in Signal Transduction.Int J Mol Sci. 2016 Oct 15;17(10):1732. doi: 10.3390/ijms17101732. Int J Mol Sci. 2016. PMID: 27754465 Free PMC article. Review.
-
Precursor of H-type II histo-blood group antigen and subterminal sialic acids on gangliosides are significantly implicated in cell entry and infection by a porcine P[11] rotavirus.Emerg Microbes Infect. 2025 Dec;14(1):2447608. doi: 10.1080/22221751.2024.2447608. Epub 2025 Jan 12. Emerg Microbes Infect. 2025. PMID: 39726161 Free PMC article.
-
Glycan Binding Specificity and Mechanism of Human and Porcine P[6]/P[19] Rotavirus VP8*s.J Virol. 2018 Jun 29;92(14):e00538-18. doi: 10.1128/JVI.00538-18. Print 2018 Jul 15. J Virol. 2018. PMID: 29720519 Free PMC article.
-
Glycan Specificity of P[19] Rotavirus and Comparison with Those of Related P Genotypes.J Virol. 2016 Oct 14;90(21):9983-9996. doi: 10.1128/JVI.01494-16. Print 2016 Nov 1. J Virol. 2016. PMID: 27558427 Free PMC article.
-
SARS-CoV-2 Receptor ACE2 Is an Interferon-Stimulated Gene in Human Airway Epithelial Cells and Is Detected in Specific Cell Subsets across Tissues.Cell. 2020 May 28;181(5):1016-1035.e19. doi: 10.1016/j.cell.2020.04.035. Epub 2020 Apr 27. Cell. 2020. PMID: 32413319 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

