Interstrand DNA-DNA cross-link formation between adenine residues and abasic sites in duplex DNA
- PMID: 24506784
- PMCID: PMC3954461
- DOI: 10.1021/ja410969x
Interstrand DNA-DNA cross-link formation between adenine residues and abasic sites in duplex DNA
Abstract
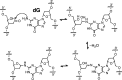
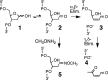
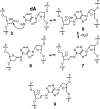
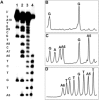
The loss of a coding nucleobase from the structure of DNA is a common event that generates an abasic (Ap) site (1). Ap sites exist as an equilibrating mixture of a cyclic hemiacetal and a ring-opened aldehyde. Aldehydes are electrophilic functional groups that can form covalent adducts with nucleophilic sites in DNA. Thus, Ap sites present a potentially reactive aldehyde as part of the internal structure of DNA. Here we report evidence that the aldehyde group of Ap sites in duplex DNA can form a covalent adduct with the N(6)-amino group of adenine residues on the opposing strand. The resulting interstrand DNA-DNA cross-link occurs at 5'-ApT/5'-AA sequences in remarkably high yields (15-70%) under physiologically relevant conditions. This naturally occurring DNA-templated reaction has the potential to generate cross-links in the genetic material of living cells.
Figures




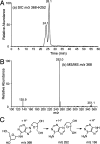

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

